 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GRMZM5G821024_P01 | ||||||||
| Common Name | LOC100285024 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Tripsacinae; Zea
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 331aa MW: 35624.6 Da PI: 6.6094 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 32.9 | 1.4e-10 | 172 | 230 | 5 | 63 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkkevaklksev 63
kr++r NR +A rs +RK +++ eLe+kv++L++e ++L +l l+ + l++e+
GRMZM5G821024_P01 172 KRAKRILANRQSAARSKERKIKYTSELERKVQTLQTEATTLSAQLTLLQRDTSGLTTEN 230
9**********************************************999888888776 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 2.5E-15 | 168 | 232 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 11.013 | 170 | 233 | IPR004827 | Basic-leucine zipper domain |
| Gene3D | G3DSA:1.20.5.170 | 1.1E-10 | 172 | 226 | No hit | No description |
| Pfam | PF00170 | 2.4E-9 | 172 | 230 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 9.85E-11 | 172 | 223 | No hit | No description |
| CDD | cd14703 | 3.47E-25 | 173 | 222 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007231 | Biological Process | osmosensory signaling pathway | ||||
| GO:0008272 | Biological Process | sulfate transport | ||||
| GO:0009294 | Biological Process | DNA mediated transformation | ||||
| GO:0009652 | Biological Process | thigmotropism | ||||
| GO:0009970 | Biological Process | cellular response to sulfate starvation | ||||
| GO:0045596 | Biological Process | negative regulation of cell differentiation | ||||
| GO:0051170 | Biological Process | nuclear import | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005829 | Cellular Component | cytosol | ||||
| GO:0003682 | Molecular Function | chromatin binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0043621 | Molecular Function | protein self-association | ||||
| GO:0051019 | Molecular Function | mitogen-activated protein kinase binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 331 aa Download sequence Send to blast |
MDLRFPPPAP SSCFGGGGVL PARGHHRRAQ SETFIRLPDA DLLLDPDADF GFSDIDFPSL 60 SDDSPAASDP TPQPQAPPPQ QPLPQAASPA SAAPPRPPSG AHMRSLSLDA AFFDGLSLQG 120 GGGVAGHKRS GSMDGATSPF EGESAPTSVL PDYAKKAVPD DRLAELALLD PKRAKRILAN 180 RQSAARSKER KIKYTSELER KVQTLQTEAT TLSAQLTLLQ RDTSGLTTEN RELKLRLQAM 240 EEQAKLRDAL NDALREEVQR LKIATGQVPN INGNPFNGGL PQQQQVPSYF SQPQGMQYFS 300 SHQGQHQHPN HHPQSSSNGG GQSLSDSMDF M |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Zm.97777 | 0.0 | cell culture| meristem| ovary| pericarp | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| Expression Atlas | GRMZM5G821024 | |||||
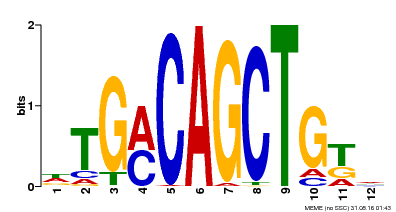
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00184 | DAP | Transfer from AT1G43700 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | GRMZM5G821024_P01 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT065699 | 0.0 | BT065699.1 Zea mays full-length cDNA clone ZM_BFc0032M22 mRNA, complete cds. | |||
| GenBank | KJ727616 | 0.0 | KJ727616.1 Zea mays clone pUT5527 bZIP transcription factor (bZIP23) mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001151391.2 | 0.0 | transcription factor RF2b | ||||
| TrEMBL | A0A1D6EHU0 | 0.0 | A0A1D6EHU0_MAIZE; Transcription factor VIP1 | ||||
| TrEMBL | A0A3L6FZV1 | 0.0 | A0A3L6FZV1_MAIZE; Transcription factor VIP1 | ||||
| STRING | GRMZM5G821024_P01 | 0.0 | (Zea mays) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1925 | 38 | 92 | Representative plant | OGRP126 | 17 | 181 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G43700.1 | 1e-50 | VIRE2-interacting protein 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | GRMZM5G821024_P01 |
| Entrez Gene | 100285024 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



