 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GRMZM2G348238_P01 | ||||||||
| Common Name | LOC107522107, Zm.30973 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Tripsacinae; Zea
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 499aa MW: 52172 Da PI: 6.1737 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 100.3 | 1.3e-31 | 274 | 329 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k+r++W+peLH+rFv+a++ LGG+++AtPk+i+elmkv+gLt+++vkSHLQkYRl+
GRMZM2G348238_P01 274 KARRCWSPELHRRFVNALQILGGAQVATPKQIRELMKVDGLTNDEVKSHLQKYRLH 329
68****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 13.69 | 271 | 331 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.69E-17 | 271 | 332 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 3.6E-28 | 272 | 332 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.9E-26 | 274 | 329 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.1E-6 | 276 | 327 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009266 | Biological Process | response to temperature stimulus | ||||
| GO:0009740 | Biological Process | gibberellic acid mediated signaling pathway | ||||
| GO:0010452 | Biological Process | histone H3-K36 methylation | ||||
| GO:0048579 | Biological Process | negative regulation of long-day photoperiodism, flowering | ||||
| GO:1903507 | Biological Process | negative regulation of nucleic acid-templated transcription | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0020127 | anatomy | primary root | ||||
| PO:0007015 | developmental stage | radicle emergence stage | ||||
| PO:0007106 | developmental stage | LP.03 three leaves visible stage | ||||
| PO:0007112 | developmental stage | 1 main shoot growth stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 499 aa Download sequence Send to blast |
MASSPSDLTL DYKPNGNGAA YAVTTPKPPQ ETLVVDGHHH HHHLTAAEQT TQKLREFLAR 60 LEEERLKIDA FKRELPLCMQ LLNHAMESYR QQLEAYQMGS LQGAPARPLV LEEFMPLKNI 120 GIDAAADKMG NPTSEKASWM ESAQLWNGPA ATADVAARGP QTPKESAECA HGAAGAGHGQ 180 RNGGGGGGAF LPFAKDKTAS AAEGAALPEL ALAPADKDAA DADRKPYLDA AAGSNNGGVL 240 GSRRDAVQSG VVNVKPAPNA PEGQQAAPPQ THRKARRCWS PELHRRFVNA LQILGGAQVA 300 TPKQIRELMK VDGLTNDEVK SHLQKYRLHT RRPMPAPPAP ATAAPQLVVL GGIWVPPEYA 360 SQAAGQAIYG AHPATQPHYT AAAAQEYYPS PAAVHHLQHH PAAAAMVHRA AAAPPPPQQA 420 AYSKAAAMAG SPPGSEGRGS AGGGSIGGGG GRERSESIEE EGEVEEREED DEDDDDDMTA 480 NKADGEGAAA GAGVGAAMY |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 4e-14 | 274 | 327 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 4e-14 | 274 | 327 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 4e-14 | 274 | 327 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 4e-14 | 274 | 327 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 4e-14 | 274 | 327 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 4e-14 | 274 | 327 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 4e-14 | 274 | 327 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 4e-14 | 274 | 327 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| Expression Atlas | GRMZM2G348238 | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may play a role in regulatory networks controlling development and metabolism. {ECO:0000250|UniProtKB:Q6Z869}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00261 | DAP | Transfer from AT2G03500 | Download |
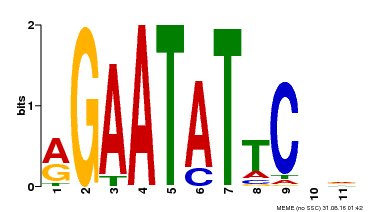
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | GRMZM2G348238_P01 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT086572 | 0.0 | BT086572.2 Zea mays full-length cDNA clone ZM_BFc0184N23 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001309287.1 | 0.0 | uncharacterized protein LOC107522107 | ||||
| Swissprot | Q5VRW2 | 1e-132 | NOH1_ORYSJ; Transcription factor NIGTH1 | ||||
| TrEMBL | A0A1D6MJX4 | 0.0 | A0A1D6MJX4_MAIZE; Myb family transcription factor EFM | ||||
| STRING | GRMZM2G348238_P01 | 0.0 | (Zea mays) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP9773 | 33 | 38 | Representative plant | OGRP4303 | 16 | 24 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G03500.1 | 9e-45 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | GRMZM2G348238_P01 |
| Entrez Gene | 107522107 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



