 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GRMZM2G341747_P01 | ||||||||
| Common Name | LOC103626260, Zm.30228 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Tripsacinae; Zea
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1020aa MW: 113248 Da PI: 5.5566 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 172 | 8.5e-54 | 21 | 138 | 2 | 118 |
CG-1 2 lke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenp 92
lke ++rwl++ ei++iL+n+++++++ e+++rp+sgsl+L++rk++ryfrkDG++w+kk d ktv+E+he+LK g+++vl+cyYah+een
GRMZM2G341747_P01 21 LKEaQHRWLRPAEICEILKNYRNFRIAPEPPNRPPSGSLFLFDRKVLRYFRKDGHNWRKKNDQKTVKEAHERLKSGSIDVLHCYYAHGEENI 112
45559*************************************************************************************** PP
CG-1 93 tfqrrcywlLeeelekivlvhylevk 118
+fqrr+yw+Lee++ +ivlvhyle+k
GRMZM2G341747_P01 113 NFQRRTYWMLEEDYMHIVLVHYLETK 138
***********************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 80.861 | 17 | 143 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 2.7E-72 | 20 | 138 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 6.8E-48 | 23 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 8.0E-6 | 446 | 533 | IPR013783 | Immunoglobulin-like fold |
| SuperFamily | SSF81296 | 2.6E-16 | 447 | 533 | IPR014756 | Immunoglobulin E-set |
| Pfam | PF01833 | 3.8E-8 | 447 | 525 | IPR002909 | IPT domain |
| SuperFamily | SSF48403 | 6.22E-17 | 640 | 743 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 4.8E-17 | 642 | 742 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.94E-13 | 642 | 739 | No hit | No description |
| PROSITE profile | PS50297 | 17.316 | 648 | 741 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 8.603 | 648 | 680 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 8.816 | 681 | 713 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.0096 | 681 | 710 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 2.39E-8 | 839 | 892 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.086 | 841 | 863 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.407 | 842 | 871 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0032 | 843 | 862 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0022 | 864 | 886 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.56 | 865 | 889 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.0E-4 | 867 | 886 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1020 aa Download sequence Send to blast |
MASAEARRLA VVPQLDIEQI LKEAQHRWLR PAEICEILKN YRNFRIAPEP PNRPPSGSLF 60 LFDRKVLRYF RKDGHNWRKK NDQKTVKEAH ERLKSGSIDV LHCYYAHGEE NINFQRRTYW 120 MLEEDYMHIV LVHYLETKGG KSSRARGNNM IQEAAVDSPL SQLPSQTMEG ESSLSGQASE 180 YEEAESDIYS GGAGHDSFTW VQQHENGTGP MIASSVFSSY TPALSIGNYH GLHATQNTSF 240 YPVNQLNSPV ILNGSSAMLG TNGCANQTDL PSWNSVIELD HEPVQMPDLQ FPVPPDQGTS 300 TEGLGVDYLT FDEVYSDGLS LQDIGATGTH GESYLQFSSG TGDLAATVNS FPQENDGSLE 360 AAIGYPFLKT QSSNLSDILK DSFKKTDSFT RWMSKELPEV EDSQIQSSSG AFWSSEEANN 420 IIEASNHEAL DQFTVSPMLS QDQLFSIVDF SPNWTYVGSK TKILVAGNIL NDSQITERSK 480 WSCMFGEVEV PANILADGTL ICYSPQHKPG RVPFYITCSN RLACSEVREF EFRPTVSQYM 540 DAPSPHGETN KVYFQIRLDK LLSLGPDEYQ ATVSNPTLEM VDLSRKISSL MASNDEWSNL 600 LKLAVDNEPS TADQQDQFAE NLIKGKLHIW LLNKVGMGGK GPSVLDDEGQ GVLHLAAALG 660 YDWAIRPTLA AGVNINFRDI HGWTALHWAA FCGRESTVVA LIALGAAPGA LTDPTPDFPG 720 STPADLASSN GQKGISGFLA ECSLTSHLQV LNLKEANMAQ ISGLPGIGDV TERDSLQPPS 780 GDSLGPVRNA TQAAARIYQV FRVQSFQRKQ AAQYEDKGGM SDERALSLLS VKPPKSGQLD 840 PLHSAATRIQ NKFRGWKGRK EFLLIRQRIV KIQAHVRGHQ VRKHYRKIVW SVGIVEKVIL 900 RWRRRGAGLR GFRSQEGSVE SSSGGTSSSS IQNKSSGDDY DFLQEGRKQT EERLQKALAR 960 VKSMAQYPEA RDQYQRIFTV VSKMQESQAM QEKMPEESAE MDMSEFKELW DDDAPIPGYF |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Zm.30228 | 0.0 | sheath | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| Expression Atlas | GRMZM2G341747 | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
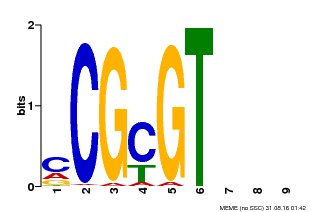
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | GRMZM2G341747_P01 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008644877.1 | 0.0 | calmodulin-binding transcription activator 3 isoform X2 | ||||
| TrEMBL | A0A1D6GRK6 | 0.0 | A0A1D6GRK6_MAIZE; Calmodulin-binding transcription activator 2 | ||||
| STRING | GRMZM2G341747_P01 | 0.0 | (Zea mays) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP608 | 38 | 140 | Representative plant | OGRP562 | 15 | 65 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | GRMZM2G341747_P01 |
| Entrez Gene | 103626260 |



