 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GRMZM2G159937_P01 | ||||||||
| Common Name | bHLH57, Zm.125686 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Tripsacinae; Zea
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 467aa MW: 49591.8 Da PI: 7.5013 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 41.5 | 2.4e-13 | 316 | 362 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCC.C...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPk.askapskKlsKaeiLekAveYIksLq 55
+h e+Er+RR+++N++f Lr ++Pk + K++Ka+ L A+ YI++L+
GRMZM2G159937_P01 316 NHVEAERQRREKLNQRFYALRAVVPKiS------KMDKASLLSDAIAYIQELE 362
799***********************66......****************996 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 8.8E-40 | 40 | 209 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 17.047 | 312 | 361 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 3.08E-15 | 315 | 366 | No hit | No description |
| Gene3D | G3DSA:4.10.280.10 | 3.1E-16 | 315 | 365 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 3.4E-16 | 315 | 366 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 7.3E-11 | 316 | 362 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 8.1E-16 | 318 | 367 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000037 | anatomy | shoot apex | ||||
| PO:0006310 | anatomy | tassel floret | ||||
| PO:0006339 | anatomy | juvenile vascular leaf | ||||
| PO:0006340 | anatomy | adult vascular leaf | ||||
| PO:0006341 | anatomy | primary shoot system | ||||
| PO:0006354 | anatomy | ear floret | ||||
| PO:0006505 | anatomy | central spike of ear inflorescence | ||||
| PO:0008018 | anatomy | transition vascular leaf | ||||
| PO:0009001 | anatomy | fruit | ||||
| PO:0009009 | anatomy | plant embryo | ||||
| PO:0009025 | anatomy | vascular leaf | ||||
| PO:0009054 | anatomy | inflorescence bract | ||||
| PO:0009066 | anatomy | anther | ||||
| PO:0009074 | anatomy | style | ||||
| PO:0009084 | anatomy | pericarp | ||||
| PO:0009089 | anatomy | endosperm | ||||
| PO:0020040 | anatomy | leaf base | ||||
| PO:0020104 | anatomy | leaf sheath | ||||
| PO:0020126 | anatomy | tassel inflorescence | ||||
| PO:0020127 | anatomy | primary root | ||||
| PO:0020136 | anatomy | ear inflorescence | ||||
| PO:0020142 | anatomy | stem internode | ||||
| PO:0020148 | anatomy | shoot apical meristem | ||||
| PO:0025142 | anatomy | leaf tip | ||||
| PO:0025287 | anatomy | seedling coleoptile | ||||
| PO:0025541 | anatomy | bundle sheath cell | ||||
| PO:0025589 | anatomy | leaf lamina tip | ||||
| PO:0001007 | developmental stage | pollen development stage | ||||
| PO:0001009 | developmental stage | D pollen mother cell meiosis stage | ||||
| PO:0001052 | developmental stage | vascular leaf expansion stage | ||||
| PO:0001053 | developmental stage | vascular leaf post-expansion stage | ||||
| PO:0001083 | developmental stage | inflorescence development stage | ||||
| PO:0001094 | developmental stage | plant embryo coleoptilar stage | ||||
| PO:0001095 | developmental stage | plant embryo true leaf formation stage | ||||
| PO:0001180 | developmental stage | plant proembryo stage | ||||
| PO:0007001 | developmental stage | early whole plant fruit ripening stage | ||||
| PO:0007003 | developmental stage | IL.03 full inflorescence length reached stage | ||||
| PO:0007006 | developmental stage | IL.00 inflorescence just visible stage | ||||
| PO:0007015 | developmental stage | radicle emergence stage | ||||
| PO:0007016 | developmental stage | whole plant flowering stage | ||||
| PO:0007022 | developmental stage | seed imbibition stage | ||||
| PO:0007026 | developmental stage | FL.00 first flower(s) open stage | ||||
| PO:0007031 | developmental stage | mid whole plant fruit ripening stage | ||||
| PO:0007032 | developmental stage | whole plant fruit formation stage up to 10% | ||||
| PO:0007045 | developmental stage | coleoptile emergence stage | ||||
| PO:0007063 | developmental stage | LP.07 seven leaves visible stage | ||||
| PO:0007065 | developmental stage | LP.05 five leaves visible stage | ||||
| PO:0007072 | developmental stage | LP.18 eighteen leaves visible stage | ||||
| PO:0007094 | developmental stage | LP.01 one leaf visible stage | ||||
| PO:0007101 | developmental stage | LP.09 nine leaves visible stage | ||||
| PO:0007104 | developmental stage | LP.15 fifteen leaves visible stage | ||||
| PO:0007106 | developmental stage | LP.03 three leaves visible stage | ||||
| PO:0007112 | developmental stage | 1 main shoot growth stage | ||||
| PO:0007116 | developmental stage | LP.11 eleven leaves visible stage | ||||
| PO:0007123 | developmental stage | LP.06 six leaves visible stage | ||||
| PO:0007633 | developmental stage | endosperm development stage | ||||
| PO:0021004 | developmental stage | inflorescence initiation stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 467 aa Download sequence Send to blast |
MAWSETDAAL FAAVLGRDAA HHLSTTPPHQ DAPAASAPEL QARLQDLVER GGAWTYGIFW 60 QESCAGGRAV LGWGDGHCRD GGAPHHDDAD RSVARKRALL RLHALYGGGD DEGADYALRL 120 DRVTAAEMYF LASMYFSFPE GAGGPGHALA TARHAWATVD PAPGWYVRAS LAQSAGLRTV 180 VFLPCKGGVL ELGSAVPVRE TPETLRALQT ALAVARPPAR EECMRIFGQD LSPGGSARAP 240 RSVDNWAPHP HLAAQATAAS ALASKEAAAG HKAPEPPRSI DFSKPGKPGH GQAGGEERRP 300 RKRGRKPANG REEPLNHVEA ERQRREKLNQ RFYALRAVVP KISKMDKASL LSDAIAYIQE 360 LEDRLRGGGG GGGGCSAARP DSPDVEVKAM QDEVVLRVTT PLYAHPVSRV FHAIRDAELI 420 VAASDVAVAD EAVTHTLVLR SPGPEQLTAE TVLAAMSRGM TSATPSP |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5gnj_A | 7e-26 | 310 | 402 | 2 | 82 | Transcription factor MYC2 |
| 5gnj_B | 7e-26 | 310 | 402 | 2 | 82 | Transcription factor MYC2 |
| 5gnj_E | 7e-26 | 310 | 402 | 2 | 82 | Transcription factor MYC2 |
| 5gnj_F | 7e-26 | 310 | 402 | 2 | 82 | Transcription factor MYC2 |
| 5gnj_G | 7e-26 | 310 | 402 | 2 | 82 | Transcription factor MYC2 |
| 5gnj_I | 7e-26 | 310 | 402 | 2 | 82 | Transcription factor MYC2 |
| 5gnj_M | 7e-26 | 310 | 402 | 2 | 82 | Transcription factor MYC2 |
| 5gnj_N | 7e-26 | 310 | 402 | 2 | 82 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 297 | 305 | RRPRKRGRK |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Zm.125686 | 0.0 | meristem | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| Expression Atlas | GRMZM2G159937 | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
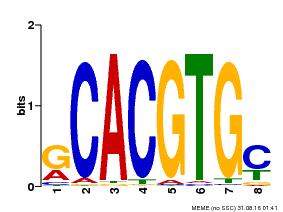
| Motif ID | Method | Source | Motif file |
| MP00099 | PBM | Transfer from AT4G16430 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | GRMZM2G159937_P01 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | EU968015 | 0.0 | EU968015.1 Zea mays clone 312846 DNA binding protein mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001150726.2 | 0.0 | uncharacterized protein LOC100284359 | ||||
| TrEMBL | A0A060D780 | 0.0 | A0A060D780_MAIZE; BHLH transcription factor (Fragment) | ||||
| STRING | GRMZM2G159937_P01 | 0.0 | (Zea mays) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP7385 | 29 | 44 | Representative plant | OGRP9430 | 10 | 12 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G16430.1 | 6e-54 | bHLH family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | GRMZM2G159937_P01 |



