 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GRMZM2G159119_P01 | ||||||||
| Common Name | LOC100216751, ZEAMMB73_380796 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Tripsacinae; Zea
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 459aa MW: 49339 Da PI: 7.061 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 101 | 7.9e-32 | 277 | 332 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k+r++W+peLH+rF++a++qLGGs++AtPk+i+elm+v+gLt+++vkSHLQkYRl+
GRMZM2G159119_P01 277 KARRCWAPELHRRFLQALQQLGGSHVATPKQIRELMNVDGLTNDEVKSHLQKYRLH 332
68****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 12.015 | 274 | 334 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.4E-28 | 275 | 335 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 5.44E-17 | 275 | 335 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 7.4E-27 | 277 | 332 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 9.7E-7 | 279 | 330 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 459 aa Download sequence Send to blast |
MEMDPPPPPP LHADRRRRFR DYLLALEEER RKIQVFQREL PLCLDLVTQT IEGMRSHMDS 60 VVVGSEETVS DHGGPVLEEF MPLKPTTLSS SSSQPHYQDE HDSAHYLEHR RGSAATANDV 120 VDADKDGEAV GDLETAAASS SRPLPHPETK KAMPDWLQSV QLWSNQQQPS ASPPQHQDEL 180 LLPCRPVALN ACRKPGGAFQ PFEKEKKKKD KEEKQRAELE LPLPAAASSA VVGDSCDRAG 240 ATDTDTDTAE NNKASSNKGG NDKEAQLSSQ SQAPSRKARR CWAPELHRRF LQALQQLGGS 300 HVATPKQIRE LMNVDGLTND EVKSHLQKYR LHTRRPNSAA AAVQSGGTSV VAPPAAPQFV 360 VVGGIWVPPP EYAAAVAVAA AAAAQPQQVH LAGDASGTTT TAADKVYAPV ATTLTPAPRP 420 RPRPRPERQS SSCSGARRSG DACSGSPAVL SSSSHTASA |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 5e-13 | 277 | 330 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 5e-13 | 277 | 330 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 5e-13 | 277 | 330 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 5e-13 | 277 | 330 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 5e-13 | 277 | 330 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 5e-13 | 277 | 330 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 5e-13 | 277 | 330 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 5e-13 | 277 | 330 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 5e-13 | 277 | 330 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 5e-13 | 277 | 330 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 5e-13 | 277 | 330 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 5e-13 | 277 | 330 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 13 | 18 | DRRRRF |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Zm.79866 | 0.0 | meristem | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| Expression Atlas | GRMZM2G159119 | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
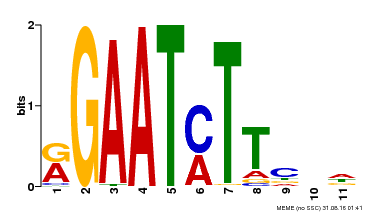
| UniProt | Transcriptional repressor that may play a role in response to nitrogen. May be involved in a time-dependent signaling for transcriptional regulation of nitrate-responsive genes. Binds specifically to the DNA sequence motif 5'-GAATC-3' or 5'-GAATATTC-3'. Represses the activity of its own promoter trough binding to these motifs. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00160 | DAP | Transfer from AT1G25550 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | GRMZM2G159119_P01 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by nitrate. {ECO:0000269|PubMed:23324170}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT037290 | 0.0 | BT037290.1 Zea mays full-length cDNA clone ZM_BFb0161K16 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001136626.1 | 0.0 | uncharacterized protein LOC100216751 | ||||
| Swissprot | Q6Z869 | 1e-112 | NIGT1_ORYSJ; Transcription factor NIGT1 | ||||
| TrEMBL | K7TM84 | 0.0 | K7TM84_MAIZE; Putative MYB DNA-binding domain superfamily protein | ||||
| STRING | GRMZM2G159119_P01 | 0.0 | (Zea mays) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6470 | 37 | 54 | Representative plant | OGRP5415 | 12 | 21 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G25550.1 | 4e-61 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | GRMZM2G159119_P01 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



