 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GRMZM2G137341_P01 | ||||||||
| Common Name | EREB16, LOC100284491 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Tripsacinae; Zea
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 255aa MW: 27241.2 Da PI: 5.8018 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 58.3 | 1.9e-18 | 79 | 129 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
+ ykGVr++ grWv+e+r+p g ++r++lg+f tae+Aa+a++ a+++l+g
GRMZM2G137341_P01 79 PVYKGVRRRN-PGRWVCEVREP--HG-KQRIWLGTFETAEMAARAHDVAALALRG 129
68*****887.9******9996..33.5*************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF00847 | 1.0E-13 | 79 | 129 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 22.089 | 80 | 137 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 1.3E-30 | 80 | 139 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 3.07E-21 | 80 | 138 | IPR016177 | DNA-binding domain |
| SMART | SM00380 | 6.0E-28 | 80 | 143 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 3.3E-8 | 81 | 92 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 7.25E-21 | 81 | 139 | No hit | No description |
| PRINTS | PR00367 | 3.3E-8 | 103 | 119 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 255 aa Download sequence Send to blast |
MLRLVPPPAR HAHGHYHSHT QAAAMDAAGS FSDYSSGTPS PVAGGGGGDD FGSSSYMTVS 60 SAPPKRRAGR TKFKETRHPV YKGVRRRNPG RWVCEVREPH GKQRIWLGTF ETAEMAARAH 120 DVAALALRGR AACLNFADSP RLLRVPPTGS GHDEIRRAAA VAADQFRPAP DQGNVAAEEE 180 AADTPPPDAL PSVTMQSVDD DPYCIIDDRL DFGMQGYLDM AQGMLIDPPP MAGSSTSGGG 240 GDDDDDDGEV KLWSY |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5wx9_A | 3e-14 | 78 | 136 | 12 | 71 | Ethylene-responsive transcription factor ERF096 |
| Search in ModeBase | ||||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| Expression Atlas | GRMZM2G137341 | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-[AG]CCGAC-3'. Binding to the C-repeat/DRE element mediates high salinity- and dehydration-inducible transcription (By similarity). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00644 | PBM | Transfer from LOC_Os02g45450 | Download |
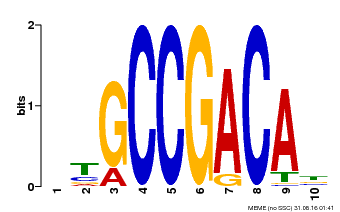
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | GRMZM2G137341_P01 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KJ727873 | 0.0 | KJ727873.1 Zea mays clone pUT5966 AP2-EREBP transcription factor (EREB16) gene, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001150858.2 | 1e-168 | dehydration-responsive element-binding protein 1A | ||||
| Swissprot | Q6EP77 | 1e-96 | DRE1G_ORYSJ; Dehydration-responsive element-binding protein 1G | ||||
| TrEMBL | A0A060D7N0 | 0.0 | A0A060D7N0_MAIZE; AP2-EREBP transcription factor (Fragment) | ||||
| STRING | GRMZM2G137341_P01 | 0.0 | (Zea mays) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP196 | 36 | 261 | Representative plant | OGRP6 | 16 | 1718 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G25470.1 | 6e-36 | C-repeat/DRE binding factor 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | GRMZM2G137341_P01 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



