 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GRMZM2G132868_P01 | ||||||||
| Common Name | LOC100283313, ZEAMMB73_451937 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Tripsacinae; Zea
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 257aa MW: 27038.3 Da PI: 7.5246 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 39.5 | 1.2e-12 | 178 | 223 | 5 | 50 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkele 50
+r++r++kNRe+A rsR RK+a+++eLe v +Le+e ++L e e
GRMZM2G132868_P01 178 QRQKRMIKNRESAARSRDRKQAYVAELESQVMQLEEEQTELLREQE 223
89*************************************9987755 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 7.5E-7 | 174 | 238 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 10.611 | 176 | 228 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF00170 | 1.6E-10 | 177 | 224 | IPR004827 | Basic-leucine zipper domain |
| CDD | cd14707 | 2.47E-17 | 178 | 223 | No hit | No description |
| SuperFamily | SSF57959 | 1.53E-10 | 179 | 227 | No hit | No description |
| Gene3D | G3DSA:1.20.5.170 | 3.2E-12 | 179 | 228 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 181 | 196 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 257 aa Download sequence Send to blast |
MASSRVMASS SPPQHAGAGS SSDLARFKSS GVGSMNMDDI IRNIYGPEAV NAAAAESAGL 60 GPAEPSLAAP EAAARRTSEE VWKEISAAGG LPAPVLHIPP AASSGAGARG GGGAAEMTLE 120 DFLARDSCGR AAVMEGNMAL GFPDGDGDAA GSVAGGVGGV RGGGKRALLD PADRAVMQRQ 180 KRMIKNRESA ARSRDRKQAY VAELESQVMQ LEEEQTELLR EQEDRRQKRL KELTERAVPV 240 IRKKSPQDLR RTNSMEW |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Zm.31952 | 0.0 | meristem| ovary| shoot| tassel | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| Expression Atlas | GRMZM2G132868 | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the ABA-responsive elements (ABREs) in vitro. Involved in abiotic stress responses and abscisic acid (ABA) signaling (PubMed:20039193). Involved in the signaling pathway that induces growth inhibition in response to D-allose (PubMed:23397192). {ECO:0000269|PubMed:20039193, ECO:0000269|PubMed:23397192}. | |||||
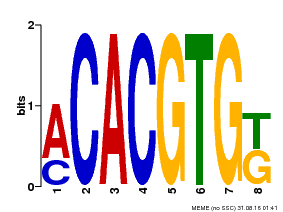
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00648 | PBM | Transfer from LOC_Os05g36160 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | GRMZM2G132868_P01 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by anoxia, drought, salt stress, oxidative stress, cold and abscisic acid (ABA) (PubMed:20039193). Induced by D-allose (PubMed:23397192). {ECO:0000269|PubMed:20039193, ECO:0000269|PubMed:23397192}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | EU964255 | 0.0 | EU964255.1 Zea mays clone 277330 G-box-binding factor 4 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001149687.1 | 0.0 | uncharacterized protein LOC100283313 | ||||
| Swissprot | Q0JHF1 | 6e-56 | BZP12_ORYSJ; bZIP transcription factor 12 | ||||
| TrEMBL | B6TGZ0 | 0.0 | B6TGZ0_MAIZE; G-box-binding factor 4 | ||||
| STRING | GRMZM2G132868_P01 | 0.0 | (Zea mays) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2665 | 37 | 86 | Representative plant | OGRP9845 | 4 | 10 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G44080.1 | 1e-15 | bZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | GRMZM2G132868_P01 |
| Entrez Gene | 100283313 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



