 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GRMZM2G102218_P01 | ||||||||
| Common Name | ZEAMMB73_586160, Zm.121737 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Tripsacinae; Zea
|
||||||||
| Family | YABBY | ||||||||
| Protein Properties | Length: 308aa MW: 33941 Da PI: 8.745 | ||||||||
| Description | YABBY family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | YABBY | 59.7 | 1.3e-18 | 104 | 148 | 2 | 46 |
YABBY 2 dvfssseqvCyvqCnfCntilavsvPstslfkvvtvrCGhCtsll 46
d++s+se++Cyv+C++Cnt+lav vP + l+ +vtv+CGhC +l
GRMZM2G102218_P01 104 DTVSQSEHLCYVRCTYCNTVLAVGVPCKRLMDTVTVKCGHCNNLS 148
78899*************************************984 PP
| |||||||
| 2 | YABBY | 93.6 | 4.9e-29 | 189 | 249 | 102 | 162 |
YABBY 102 senedeevprvppvirPPekrqrvPsaynrfikeeiqrikasnPdishreafsaaaknWah 162
s +++ +pr+p v++PPek+ r Psaynrf++eeiqrika+ Pdi hreafs+aaknWa
GRMZM2G102218_P01 189 SPTSTDLTPRMPFVVKPPEKKHRLPSAYNRFMREEIQRIKAAKPDIPHREAFSMAAKNWAK 249
556677789**************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF04690 | 7.3E-46 | 108 | 250 | IPR006780 | YABBY protein |
| SuperFamily | SSF47095 | 6.15E-8 | 197 | 251 | IPR009071 | High mobility group box domain |
| Gene3D | G3DSA:1.10.30.10 | 4.1E-5 | 204 | 251 | IPR009071 | High mobility group box domain |
| CDD | cd01390 | 4.22E-5 | 211 | 252 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010254 | Biological Process | nectary development | ||||
| GO:0010582 | Biological Process | floral meristem determinacy | ||||
| GO:0048479 | Biological Process | style development | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 308 aa Download sequence Send to blast |
MCAWGPGIIA GIETLPPSPL VACSTDSSSS SSTSRPVSSP FFVCLCLLVS RLTTQLASRQ 60 PDTRCFQISH RTVVVGPSEN IPPRPPPRAP PRPRTPAGHQ ASMDTVSQSE HLCYVRCTYC 120 NTVLAVGVPC KRLMDTVTVK CGHCNNLSYL SPRPPMVQPL SPTDHPLGPF QCQGPCSECR 180 RNQPLPLASP TSTDLTPRMP FVVKPPEKKH RLPSAYNRFM REEIQRIKAA KPDIPHREAF 240 SMAAKNWAKC DPRCSTTAST ATSNSAPEPA RVVVPTPHVT EPRFDLEDRA KEHVIESFDI 300 FKQIERNI |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Zm.121737 | 0.0 | ear| endosperm| silk | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| Expression Atlas | GRMZM2G102218 | |||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Detected during flower and leaf development. Expression in the flower meristem in the early stage of flower development. When carpel primordia begin to form, specific and uniform expression in carpel primordia. Expression in the central region of the leaf plastochron 1 (P1) primordia. Detected up to P4 stage, hardly detected in the P5 leaves. {ECO:0000269|PubMed:14729915}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Regulates carpel specification in flower development. Severe or intermediate mutation in DL causes complete or partial homeotic conversion of carpels into stamens without affecting the identities of other floral organs. Interacts antagonistically with class B genes and controls floral meristem determinacy. Regulates midrib formation in leaves probably by inducing cell proliferation in the central region of the leaf. {ECO:0000269|PubMed:14729915}. | |||||
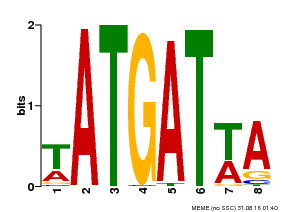
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00220 | DAP | Transfer from AT1G69180 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | GRMZM2G102218_P01 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT087325 | 0.0 | BT087325.1 Zea mays full-length cDNA clone ZM_BFb0040H18 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001183591.1 | 1e-150 | uncharacterized protein LOC100502185 | ||||
| Swissprot | Q76EJ0 | 1e-110 | YABDL_ORYSJ; Protein DROOPING LEAF | ||||
| TrEMBL | C4J928 | 1e-149 | C4J928_MAIZE; DL related protein | ||||
| STRING | GRMZM2G102218_P01 | 0.0 | (Zea mays) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5951 | 36 | 57 | Representative plant | OGRP9069 | 11 | 14 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G69180.1 | 5e-50 | YABBY family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | GRMZM2G102218_P01 |



