 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GRMZM2G076636_P02 | ||||||||
| Common Name | cl6102_1a, ZEAMMB73_729450 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Tripsacinae; Zea
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 611aa MW: 66406.8 Da PI: 7.3115 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 39.3 | 1.1e-12 | 457 | 503 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCC.C...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPk.askapskKlsKaeiLekAveYIksLq 55
+h e+Er+RR+++N++f Lr ++P+ + K++Ka+ L A+ YI +Lq
GRMZM2G076636_P02 457 NHVEAERQRREKLNQRFYALRAVVPNiS------KMDKASLLGDAITYITDLQ 503
799***********************66......*****************99 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 1.4E-51 | 60 | 245 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 16.831 | 453 | 502 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 4.70E-15 | 456 | 507 | No hit | No description |
| SuperFamily | SSF47459 | 6.15E-19 | 456 | 520 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 3.7E-10 | 457 | 503 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 2.8E-18 | 457 | 516 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.9E-16 | 459 | 508 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 611 aa Download sequence Send to blast |
MVMKMEHEDN GAIGGTDGTW TEDDRALGAA VLGADAFAYL TKGGGAISEG LVATSLPGDL 60 QNKLQELVES ESPGTSWNYA IFWQLSRTKS GDLVLGWGDG CCREPRDGEL GAAASAGSED 120 SKQRMRKRAL QRLHIAFGVA DEEDYSPGID QVTDTEMFFL ASMYFAFPRH AGGPGQAFAA 180 GIPIWVPNSE RKVVPANYCY RGFLANAAGF RTIVLVPFES GVLELGSTQH IAESSGTVQT 240 VRSVFAGTSG NKSAVQRHEA ERSPGLAKIF GKDLNLGRPS VGLAVGVSNS KVDERTWEQR 300 SAVGGTSLLP SVQKGLQNFS WSQARGLNSH QQKFGNGVLI VSNNEGAHRN NGAVDSPSAA 360 QFQLQKAPQL QKLSVVQKTP QLVNQQPMQA QVPRQIDFSA GSSSKPGVLV TRAGVLDGES 420 AEVDGLCKEE GPPPVMEDRR PRKRGRKPAN GREEPLNHVE AERQRREKLN QRFYALRAVV 480 PNISKMDKAS LLGDAITYIT DLQKKLKEME TERERLLESG MVDPRERAPR PEVDIQVVQD 540 EVLVRVMSPM ENHPVKKVFQ AFEEAEVRVG ESKVTSNNNG TAVHSFIIKC PGTEQQTREK 600 VIAAMSRAMS S |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5gnj_A | 6e-28 | 451 | 513 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_B | 6e-28 | 451 | 513 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_E | 6e-28 | 451 | 513 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_F | 6e-28 | 451 | 513 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_G | 6e-28 | 451 | 513 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_I | 6e-28 | 451 | 513 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_M | 6e-28 | 451 | 513 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_N | 6e-28 | 451 | 513 | 2 | 64 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 438 | 446 | RRPRKRGRK |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Zm.25463 | 0.0 | endosperm| meristem | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| Expression Atlas | GRMZM2G076636 | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
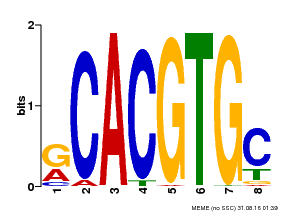
| Motif ID | Method | Source | Motif file |
| MP00100 | PBM | Transfer from AT1G01260 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | GRMZM2G076636_P02 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT053968 | 0.0 | BT053968.1 Zea mays full-length cDNA clone ZM_BFc0083E19 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001145780.2 | 0.0 | uncharacterized protein LOC100279287 | ||||
| Refseq | XP_020397645.1 | 0.0 | uncharacterized protein LOC100279287 isoform X1 | ||||
| Refseq | XP_020397646.1 | 0.0 | uncharacterized protein LOC100279287 isoform X1 | ||||
| TrEMBL | K7VBN7 | 0.0 | K7VBN7_MAIZE; Putative HLH DNA-binding domain superfamily protein | ||||
| STRING | GRMZM2G076636_P01 | 0.0 | (Zea mays) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G01260.3 | 1e-131 | bHLH family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | GRMZM2G076636_P02 |



