 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GRMZM2G065374_P01 | ||||||||
| Common Name | Zm.6794 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Tripsacinae; Zea
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 523aa MW: 56431.2 Da PI: 6.8215 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 53.4 | 4.6e-17 | 328 | 374 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
hn ErrRRdriN+++ L+el+P++ K +Ka+iL +++eY+ksLq
GRMZM2G065374_P01 328 VHNLSERRRRDRINEKMRALQELIPHC-----NKTDKASILDETIEYLKSLQ 374
6*************************8.....6******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00083 | 2.15E-17 | 320 | 378 | No hit | No description |
| SuperFamily | SSF47459 | 8.9E-21 | 322 | 388 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 18.482 | 324 | 373 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 1.5E-21 | 328 | 383 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 2.6E-14 | 328 | 374 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.3E-17 | 330 | 379 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0009693 | Biological Process | ethylene biosynthetic process | ||||
| GO:0010244 | Biological Process | response to low fluence blue light stimulus by blue low-fluence system | ||||
| GO:0010600 | Biological Process | regulation of auxin biosynthetic process | ||||
| GO:0010928 | Biological Process | regulation of auxin mediated signaling pathway | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016021 | Cellular Component | integral component of membrane | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0009001 | anatomy | fruit | ||||
| PO:0009089 | anatomy | endosperm | ||||
| PO:0020127 | anatomy | primary root | ||||
| PO:0020142 | anatomy | stem internode | ||||
| PO:0020148 | anatomy | shoot apical meristem | ||||
| PO:0025287 | anatomy | seedling coleoptile | ||||
| PO:0001095 | developmental stage | plant embryo true leaf formation stage | ||||
| PO:0007015 | developmental stage | radicle emergence stage | ||||
| PO:0007031 | developmental stage | mid whole plant fruit ripening stage | ||||
| PO:0007045 | developmental stage | coleoptile emergence stage | ||||
| PO:0007106 | developmental stage | LP.03 three leaves visible stage | ||||
| PO:0007112 | developmental stage | 1 main shoot growth stage | ||||
| PO:0007633 | developmental stage | endosperm development stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 523 aa Download sequence Send to blast |
MSRVNGVTDA LSIGPSAGYI LFLVIVHILL FFTAYDRSYT LLAIQKVTLL CSACKAPRTD 60 GGDDSQGSSA RARQQQHAAV LRSESGEKTL FSGHSSRSPD EMDGNARSAA TNQKKPVVAD 120 DDLVELLWHN GSVVAQPQAH QRPAPTYDPD RPGTSGLTGE ETAAWFPDTL DDALEKDLYT 180 QLWYSTIADA APQHEGTLPG PTSQPSPSPP VGSSGVESSW AGDICSTFCG SNQVLRTPAR 240 IRGKDAALQS ELPSNTGAHD GTSSSGGSGS NYGGFGLPSD SVHVQKRKGR CRDDSDSPSE 300 DAECEEASEE TKPSRRYGTK RRTRAAEVHN LSERRRRDRI NEKMRALQEL IPHCNKTDKA 360 SILDETIEYL KSLQMQVQIM WMTSGMAPMM FPGVHQFIPQ MALGMNPGCI PAAQGLSQMP 420 RLPYMNHTLP NHIHLNSSPA MNLMNPLNAA NQVQIGHLRD ASSHLLHLDG GRAAVVPQVP 480 GPGPHVHGHQ IAQAEEHNKI LEVAASTVIP TSKAGQPPTL HRV |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 319 | 336 | KRRTRAAEVHNLSERRRR |
| 2 | 332 | 337 | ERRRRD |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Zm.37440 | 0.0 | meristem | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| Expression Atlas | GRMZM2G065374 | |||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Highly expressed in the node portions of the stem. Expressed in the leaves and the basal part of shoots. {ECO:0000269|PubMed:22984180}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that may act as negative regulator of phyB-dependent light signal transduction (PubMed:17485859). Transcription activator that acts as positive regulator of internode elongation. May function via regulation of cell wall-related genes. May play a role in a drought-associated growth-restriction mechanism in response to drought stress (PubMed:22984180). {ECO:0000269|PubMed:17485859, ECO:0000269|PubMed:22984180}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00082 | ChIP-seq | Transfer from AT3G59060 | Download |
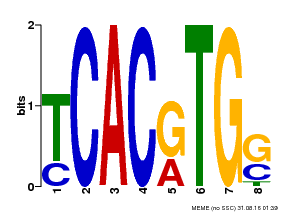
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | GRMZM2G065374_P01 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by light in dark-grown etiolated seedlings (PubMed:17485859). Circadian oscillation under 12 h light/12 h dark cycle conditions, with peaks in the middle of the light period (PubMed:17485859, PubMed:22984180). Down-regulated by cold and drought stresses (PubMed:22984180). {ECO:0000269|PubMed:17485859, ECO:0000269|PubMed:22984180}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT069545 | 1e-148 | BT069545.1 Zea mays full-length cDNA clone ZM_BFb0157L03 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008665523.1 | 0.0 | transcription factor PIF5 isoform X1 | ||||
| Refseq | XP_008665524.1 | 0.0 | transcription factor PIF5 isoform X1 | ||||
| Refseq | XP_008665525.1 | 0.0 | transcription factor PIF5 isoform X1 | ||||
| Refseq | XP_008665526.1 | 0.0 | transcription factor PIF5 isoform X1 | ||||
| Refseq | XP_008665527.1 | 0.0 | transcription factor PIF5 isoform X1 | ||||
| Refseq | XP_020402574.1 | 0.0 | transcription factor PIF5 isoform X1 | ||||
| Swissprot | Q10CH5 | 1e-153 | PIL13_ORYSJ; Transcription factor PHYTOCHROME INTERACTING FACTOR-LIKE 13 | ||||
| TrEMBL | A0A1D6L6L0 | 0.0 | A0A1D6L6L0_MAIZE; Transcription factor PIF4 | ||||
| STRING | GRMZM2G065374_P01 | 0.0 | (Zea mays) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3174 | 34 | 82 | Representative plant | OGRP258 | 16 | 128 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G59060.1 | 4e-36 | phytochrome interacting factor 3-like 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | GRMZM2G065374_P01 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



