 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GRMZM2G061734_P01 | ||||||||
| Common Name | LOC100279240 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Tripsacinae; Zea
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 429aa MW: 44959.3 Da PI: 7.1839 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 130.7 | 5.1e-41 | 109 | 185 | 2 | 78 |
-SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 2 CqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
C v+gC+adls++++yhrrhkvCe+hsk+pvv+v+g+e rfCqqCsrfh l efDe+krsCr+rL++hn+rrrk+q+
GRMZM2G061734_P01 109 CAVDGCKADLSKCRDYHRRHKVCEAHSKTPVVVVAGREMRFCQQCSRFHLLLEFDEAKRSCRKRLDGHNRRRRKPQP 185
**************************************************************************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 1.1E-35 | 101 | 170 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.163 | 106 | 183 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.03E-38 | 107 | 188 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 1.1E-30 | 109 | 182 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0048653 | Biological Process | anther development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 429 aa Download sequence Send to blast |
MDWDLKMPVS WDLPDLEHDA MPPPPPVSAA STAAASGIAA AAAAPSSATA PSRAECSVDL 60 KLGGLGEFGA ADGTATKEPA AATAAPSASP MKRPRLGPGG AGGAQCPSCA VDGCKADLSK 120 CRDYHRRHKV CEAHSKTPVV VVAGREMRFC QQCSRFHLLL EFDEAKRSCR KRLDGHNRRR 180 RKPQPDTMNS GSFMTSQQGL FSSAGTRFSS FPAPRPEPSW SGVIKSEDSS YYTHHPVLSN 240 RPHVAGTSTS PAYSKEGRRF PFLQDGDQVS FSASGAGTLE VSTVCQPLLK TTAAVAPPPP 300 ESSKMLAPVL DSDCALSLLS SPANSSSVDV SRMVQPAERI PMAQPLVPGL QHHQFGGSPV 360 PDWFAGSGAV PAAGTGGFAC PHSVESEQFN TVLVPSSDGG HEMNYHGIFH VGGEGSSDGT 420 SPSLPFSWQ |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 1e-27 | 99 | 182 | 1 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 165 | 182 | KRSCRKRLDGHNRRRRKP |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Zm.18751 | 0.0 | meristem| shoot | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| Expression Atlas | GRMZM2G061734 | |||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in young panicles. {ECO:0000269|PubMed:16861571}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' (By similarity). May be involved in panicle development. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00555 | DAP | Transfer from AT5G50570 | Download |
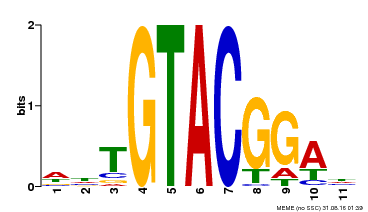
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | GRMZM2G061734_P01 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Negatively regulated by microRNAs miR156b and miR156h. {ECO:0000305|PubMed:16861571}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT053875 | 0.0 | BT053875.1 Zea mays full-length cDNA clone ZM_BFb0379K14 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001145733.2 | 0.0 | SBP-transcription factor 5 | ||||
| Swissprot | Q0J0K1 | 1e-154 | SPL18_ORYSJ; Squamosa promoter-binding-like protein 18 | ||||
| TrEMBL | B7ZX33 | 0.0 | B7ZX33_MAIZE; Uncharacterized protein | ||||
| STRING | GRMZM2G061734_P01 | 0.0 | (Zea mays) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1755 | 37 | 95 | Representative plant | OGRP97 | 17 | 230 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G50670.1 | 9e-44 | SBP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | GRMZM2G061734_P01 |



