 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GRMZM2G017087_P03 | ||||||||
| Common Name | kn1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Tripsacinae; Zea
|
||||||||
| Family | TALE | ||||||||
| Protein Properties | Length: 193aa MW: 22580.5 Da PI: 7.4198 | ||||||||
| Description | TALE family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 28.8 | 2.1e-09 | 122 | 156 | 21 | 55 |
HSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHH CS
Homeobox 21 knrypsaeereeLAkklgLterqVkvWFqNrRake 55
k +yps++++ LA+++gL+ +q+ +WF N+R ++
GRMZM2G017087_P03 122 KWPYPSETQKVALAESTGLDLKQINNWFINQRKRH 156
569*****************************885 PP
| |||||||
| 2 | ELK | 38.2 | 3e-13 | 76 | 97 | 1 | 22 |
ELK 1 ELKhqLlrKYsgyLgsLkqEFs 22
ELKh+Ll+KYsgyL+sLkqE+s
GRMZM2G017087_P03 76 ELKHHLLKKYSGYLSSLKQELS 97
9*******************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03791 | 3.9E-16 | 1 | 36 | IPR005541 | KNOX2 |
| SMART | SM01256 | 3.6E-13 | 1 | 38 | IPR005541 | KNOX2 |
| Pfam | PF03789 | 3.6E-10 | 76 | 97 | IPR005539 | ELK domain |
| PROSITE profile | PS51213 | 11.331 | 76 | 96 | IPR005539 | ELK domain |
| SMART | SM01188 | 4.9E-7 | 76 | 97 | IPR005539 | ELK domain |
| PROSITE profile | PS50071 | 12.698 | 96 | 159 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 2.27E-20 | 97 | 171 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 1.6E-12 | 98 | 163 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 4.4E-29 | 101 | 161 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 1.90E-11 | 108 | 160 | No hit | No description |
| Pfam | PF05920 | 4.8E-17 | 116 | 155 | IPR008422 | Homeobox KN domain |
| PROSITE pattern | PS00027 | 0 | 134 | 157 | IPR017970 | Homeobox, conserved site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 193 aa Download sequence Send to blast |
MEAYHEMLVK FREELTRPLQ EAMEFMRRVE SQLNSLSISG RSLRNILSSG SSEEDQEGSG 60 GETELPEVDA HGVDQELKHH LLKKYSGYLS SLKQELSKKK KKGKLPKEAR QQLLSWWDQH 120 YKWPYPSETQ KVALAESTGL DLKQINNWFI NQRKRHWKPS EEMHHLMMDG YHTTNAFYMD 180 GHFINDGGLY RLG |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Zm.94710 | 0.0 | ear| meristem| ovary| shoot| tassel | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| Expression Atlas | GRMZM2G017087 | |||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Expressed throughout apical and vegetative meristem during development. Down-regulated as leaves and floral organs are initiated. {ECO:0000269|PubMed:1362381}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in apical meristems of vegetative and floral stems as well as in the underlying ground meristem. Specifically expressed in vascular bundles developing both in the leaf and stem. Very low levels of expression in leaves. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds to RNA (PubMed:17965274). Possible transcription factor that regulates genes involved in development. Mutations in KN-1 alter leaf development. Foci of cells along the lateral vein do not differentiate properly but continue to divide, forming knots. May participate in the switch from indeterminate to determinate cell fates. Probably binds to the DNA sequence 5'-TGAC-3'. {ECO:0000269|PubMed:17965274}. | |||||
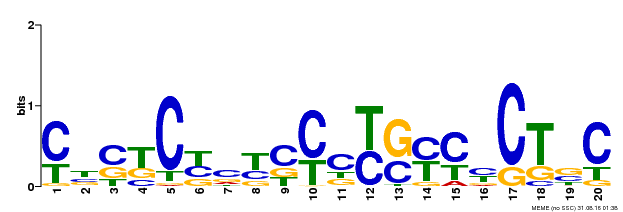
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00675 | ChIP-seq | SRX158000 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | GRMZM2G017087_P03 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | EU961952 | 0.0 | EU961952.1 Zea mays clone 239250 homeobox protein OSH1 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001266662.2 | 1e-140 | homeotic protein knotted-1 | ||||
| Swissprot | P24345 | 1e-141 | KN1_MAIZE; Homeotic protein knotted-1 | ||||
| TrEMBL | A0A1D6L2Y0 | 1e-141 | A0A1D6L2Y0_MAIZE; Homeotic protein knotted-1 | ||||
| STRING | GRMZM2G017087_P02 | 1e-140 | (Zea mays) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G08150.1 | 3e-60 | KNOTTED-like from Arabidopsis thaliana | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | GRMZM2G017087_P03 |



