 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_015899050.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rhamnaceae; Paliureae; Ziziphus
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 403aa MW: 44024.4 Da PI: 7.6816 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 39.8 | 7.9e-13 | 331 | 376 | 6 | 55 |
HHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 6 nerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
+++Er RR ri +++ +L+el+P+ +k + a++L +AveYIk+Lq
XP_015899050.1 331 SIAERVRRTRISERMRKLQELVPNM----DKQTNTADMLDLAVEYIKDLQ 376
689*********************7....688*****************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50888 | 15.148 | 325 | 375 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 7.33E-16 | 328 | 389 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 1.74E-12 | 329 | 378 | No hit | No description |
| Gene3D | G3DSA:4.10.280.10 | 1.7E-14 | 330 | 386 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 8.4E-10 | 331 | 376 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.0E-11 | 331 | 381 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 403 aa Download sequence Send to blast |
MDSNTHQSFH HQQQQDHQSN SGLLRFRSAP SSLLINFTEE GGGDCGGGVN KNGSCEGSES 60 ERLFTRFVNY SGNNDSDSTP SFQDFDDKST VTATEAAVVS NRMNSQQQGY NTGLPPHYPR 120 HGSNTSASAM DNSSFGLVGS MAMDHQAQSK TVHSNLARQN STPAGLFSHI SVPNGNYGGL 180 NGSNGEVSPP TNRLKSHLSF SSRFPTSLGM LSQISEIGSE NLGAGGLEEG KLGSSNGDGR 240 FYGTGFPLGS WSDSSNFTES LGGLRRDQDN DRKLFSSTQN EEFGNRIHLL SHHLSLPKTS 300 ADVAMEKLLH FQDSVPCKIR AKRGCATHPR SIAERVRRTR ISERMRKLQE LVPNMDKQTN 360 TADMLDLAVE YIKDLQKQFK TLSDNRANCK CLNIQKPVSN QVV |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00308 | DAP | Transfer from AT2G42280 | Download |
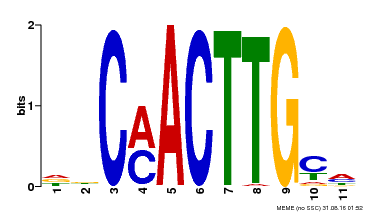
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_015899050.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015899050.1 | 0.0 | transcription factor bHLH130 isoform X2 | ||||
| Swissprot | Q66GR3 | 1e-102 | BH130_ARATH; Transcription factor bHLH130 | ||||
| TrEMBL | A0A2P5F1I4 | 0.0 | A0A2P5F1I4_TREOI; Basic helix-loop-helix transcription factor | ||||
| STRING | XP_010111430.1 | 1e-166 | (Morus notabilis) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G42280.1 | 4e-99 | bHLH family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



