 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_015895933.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rhamnaceae; Paliureae; Ziziphus
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 832aa MW: 93690.9 Da PI: 6.7169 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 71.7 | 9.3e-23 | 133 | 234 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrse 89
f+k+lt sd++++g +++ k++a+e+ + k++ +++l+ +d++g++W +++i+r++++r++l++GW+ Fv++++L +gD ++F + ++
XP_015895933.1 133 FCKTLTASDTSTHGGFSVLKRHADEClprldmS-KQPPNQELVAKDLHGNEWHFRHIFRGHPKRHLLQSGWSLFVSSKKLVAGDAFIFL--RGET 224
99*****************************63.4445569************************************************..3488 PP
E..EEEEE-S CS
B3 90 felvvkvfrk 99
+e +v+v+r+
XP_015895933.1 225 GEHRVGVRRA 234
8899**9986 PP
| |||||||
| 2 | Auxin_resp | 118.8 | 4.3e-39 | 259 | 341 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
a+ha+st+++F+v+Y+Pr+s++eF+v+++++++++k+++ +GmRfkm+fe+e+++e+r+sGtvvg +d+dpvrWp+SkWr+Lk
XP_015895933.1 259 AWHAVSTGTMFTVYYKPRTSPAEFIVPFDEYMESVKNSYPIGMRFKMRFEGEEAPEQRFSGTVVGAEDADPVRWPGSKWRCLK 341
89*******************************************************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 1.6E-39 | 125 | 248 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 4.51E-39 | 127 | 262 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 1.04E-19 | 131 | 233 | No hit | No description |
| Pfam | PF02362 | 1.8E-20 | 133 | 234 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 11.265 | 133 | 235 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 6.3E-20 | 133 | 235 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 3.0E-36 | 259 | 341 | IPR010525 | Auxin response factor |
| Pfam | PF02309 | 2.0E-10 | 703 | 789 | IPR033389 | AUX/IAA domain |
| SuperFamily | SSF54277 | 3.6E-12 | 703 | 786 | No hit | No description |
| PROSITE profile | PS51745 | 24.11 | 707 | 791 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008285 | Biological Process | negative regulation of cell proliferation | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010047 | Biological Process | fruit dehiscence | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0010227 | Biological Process | floral organ abscission | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 832 aa Download sequence Send to blast |
MEYNTGNNNT SSRASATFPV ADHEDALYKE LWHACAGPLV TVPRQGQLVF YFPQGHIEQV 60 EASTNQVVDQ QMPAYDLPSK ILCRVINVLL KAETDTDEVF AQVTLLPEPK QDENLVVKET 120 PPPPSSRPRV HSFCKTLTAS DTSTHGGFSV LKRHADECLP RLDMSKQPPN QELVAKDLHG 180 NEWHFRHIFR GHPKRHLLQS GWSLFVSSKK LVAGDAFIFL RGETGEHRVG VRRAMRQMSN 240 IPSSVISSHS MHIGVLATAW HAVSTGTMFT VYYKPRTSPA EFIVPFDEYM ESVKNSYPIG 300 MRFKMRFEGE EAPEQRFSGT VVGAEDADPV RWPGSKWRCL KVRWDETSPV HRPERVSPWK 360 IEPALAHAPD HLPACQMKRP RVSVPSSTNY SQHAIEGSSK INVETSTRNE FSMALQGQEI 420 LTPRGTFGRN DERDTAQNVN LWTASQGPDQ AEPNFGNTVF GTEDRVSQTR HRTHCMNQIF 480 GSRSLQESYR VGWPLVDQNP VNVSQLRKQV SDQEGIYNLS ASSESTMHSP SSNMMESNMK 540 LSVSEVSGLQ YHKPANSRYR AHSEYDGLHG VEVKQQPGHW LLPLLPPSYP ENSNNPVALN 600 LDHHHMQHEE KGKGTGNCKL FGISLISGPD VTEQDMLNKN SMHRPQVQAD VTTEHHQDMG 660 SEQVLEQLNY SKSAETAIRG DDAGKPFQVL KQRYLDSQVK HQGGSTRSCI KVHKQGTALG 720 RSVDLSKFNG YDELISELDR IFEFNGELNS PKKKWLIVFT DNEGDMMLVG DDPWQEFCHI 780 VRKITIYTRE DVKRMDLQTT KLEEISATAD PRLGSEEDRS LDHPSAASPY EG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldv_A | 1e-170 | 25 | 363 | 17 | 354 | Auxin response factor 1 |
| 4ldw_A | 1e-170 | 25 | 363 | 17 | 354 | Auxin response factor 1 |
| 4ldw_B | 1e-170 | 25 | 363 | 17 | 354 | Auxin response factor 1 |
| 4ldx_A | 1e-170 | 25 | 363 | 17 | 354 | Auxin response factor 1 |
| 4ldx_B | 1e-170 | 25 | 363 | 17 | 354 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). Could act as transcriptional activator or repressor. Formation of heterodimers with Aux/IAA proteins may alter their ability to modulate early auxin response genes expression. Promotes flowering, stamen development, floral organ abscission and fruit dehiscence. Functions independently of ethylene and cytokinin response pathways. May act as a repressor of cell division and organ growth. {ECO:0000269|PubMed:12036261, ECO:0000269|PubMed:15960614, ECO:0000269|PubMed:16176952, ECO:0000269|PubMed:16339187}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00574 | DAP | Transfer from AT5G62000 | Download |
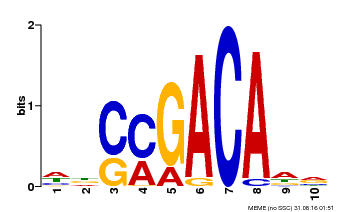
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_015895933.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015895933.1 | 0.0 | auxin response factor 2-like | ||||
| Swissprot | Q94JM3 | 0.0 | ARFB_ARATH; Auxin response factor 2 | ||||
| TrEMBL | A0A2P5F1Y5 | 0.0 | A0A2P5F1Y5_TREOI; Auxin response factor | ||||
| STRING | XP_010100039.1 | 0.0 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3281 | 33 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G62000.3 | 0.0 | auxin response factor 2 | ||||



