 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_015890577.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rhamnaceae; Paliureae; Ziziphus
|
||||||||
| Family | AP2 | ||||||||
| Protein Properties | Length: 398aa MW: 44599.3 Da PI: 7.3568 | ||||||||
| Description | AP2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 48 | 3e-15 | 232 | 291 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrd.pse.ng..kr.krfslgkfgtaeeAakaaiaarkkleg 55
s y+GV++++++gr++A+++d + g ++ ++++lg ++ +e+Aa+a++ a++k++g
XP_015890577.1 232 SIYRGVTRHRWTGRYEAHLWDnSCRrEGqaRKgRQVYLGGYDKEEKAARAYDLAALKYWG 291
57*******************666664478446*************************98 PP
| |||||||
| 2 | AP2 | 27.3 | 8.9e-09 | 334 | 368 | 1 | 38 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgt 38
s y+GV++++++grW A+I + +k +lg+fg
XP_015890577.1 334 SIYRGVTRHHQQGRWQARIGRVAG---NKDLYLGTFGK 368
57****************988532...5*********7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF00847 | 4.8E-12 | 232 | 291 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 8.5E-18 | 232 | 300 | IPR016177 | DNA-binding domain |
| CDD | cd00018 | 9.08E-23 | 232 | 301 | No hit | No description |
| PROSITE profile | PS51032 | 19.44 | 233 | 299 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 1.8E-16 | 233 | 300 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 4.0E-29 | 233 | 305 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 7.4E-6 | 234 | 245 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 7.4E-6 | 281 | 301 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 3.1E-4 | 334 | 368 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 5.04E-6 | 334 | 370 | IPR016177 | DNA-binding domain |
| PROSITE profile | PS51032 | 8.887 | 335 | 394 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 3.7E-4 | 335 | 367 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 0.0014 | 335 | 396 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010080 | Biological Process | regulation of floral meristem growth | ||||
| GO:0010492 | Biological Process | maintenance of shoot apical meristem identity | ||||
| GO:0035265 | Biological Process | organ growth | ||||
| GO:0048364 | Biological Process | root development | ||||
| GO:0060772 | Biological Process | leaf phyllotactic patterning | ||||
| GO:0060774 | Biological Process | auxin mediated signaling pathway involved in phyllotactic patterning | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 398 aa Download sequence Send to blast |
MAPATNWLSF SLSPMEMLRS SESTQFMAYD NSNTVSPQYL IDNFYANGWT NPKSQMYFPE 60 EDNNQGKECQ GPSNIAADSP IITSFIDPQT HNQAIPKLED FLGDSSSMVR YSDSQTETQD 120 SSLTQIYDQG SAYFNDQQQD LKAIAAFQTF SANSGSEVDD SVSIARTQQL TGGEFTTGHS 180 IDSSGAGNEL GFSSCPTGSL SLAVTQSSEK AIVSADSDCT KKIADTFGQR TSIYRGVTRH 240 RWTGRYEAHL WDNSCRREGQ ARKGRQVYLG GYDKEEKAAR AYDLAALKYW GPTATTNFPV 300 TNYTKELEEM KHVTKQEFIA SLRRKSSGFS RGASIYRGVT RHHQQGRWQA RIGRVAGNKD 360 LYLGTFGKFR LLSYMFAAII FWLKGHYKHH LVWEQSSH |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probably acts as a transcriptional activator. Binds to the GCC-box pathogenesis-related promoter element. May be involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways (By similarity). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00504 | DAP | Transfer from AT5G10510 | Download |
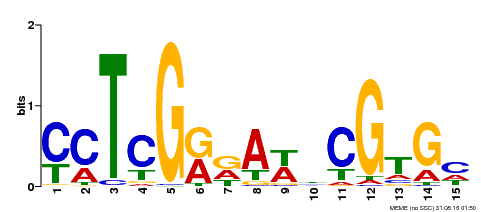
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_015890577.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015890577.1 | 0.0 | AP2-like ethylene-responsive transcription factor AIL6 | ||||
| Swissprot | Q52QU2 | 1e-160 | AIL6_ARATH; AP2-like ethylene-responsive transcription factor AIL6 | ||||
| TrEMBL | A0A2P5EGR7 | 0.0 | A0A2P5EGR7_TREOI; AP2/ERF transcription factor | ||||
| STRING | XP_010113448.1 | 0.0 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF4810 | 31 | 49 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G10510.2 | 1e-163 | AINTEGUMENTA-like 6 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



