 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_015886300.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rhamnaceae; Paliureae; Ziziphus
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1057aa MW: 118383 Da PI: 5.3202 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 185.6 | 5.2e-58 | 12 | 127 | 3 | 118 |
CG-1 3 kekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrr 97
+ ++rwl++ ei++iL n++k+++t+e++trp+sgsl+L++rk++ryfrkDG++w+kkkdgktv+E+hekLKvg+++vl+cyYah+e+n++fqrr
XP_015886300.1 12 EAQHRWLRPAEICEILRNYQKFRITSEPPTRPPSGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHEKLKVGSIDVLHCYYAHGEDNENFQRR 106
449******************************************************************************************** PP
CG-1 98 cywlLeeelekivlvhylevk 118
+yw+Le+el +iv+vhylevk
XP_015886300.1 107 SYWMLEQELMHIVFVHYLEVK 127
******************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 85.308 | 6 | 132 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 2.2E-84 | 9 | 127 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 1.5E-51 | 12 | 126 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 3.4E-5 | 445 | 556 | IPR013783 | Immunoglobulin-like fold |
| SuperFamily | SSF81296 | 3.67E-14 | 471 | 556 | IPR014756 | Immunoglobulin E-set |
| Pfam | PF01833 | 1.7E-4 | 471 | 546 | IPR002909 | IPT domain |
| PROSITE profile | PS50297 | 19.571 | 659 | 766 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 2.64E-18 | 659 | 766 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 4.10E-16 | 661 | 764 | No hit | No description |
| Gene3D | G3DSA:1.25.40.20 | 3.6E-18 | 663 | 770 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 6.9E-8 | 666 | 731 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 9.511 | 672 | 704 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 670 | 672 | 701 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 10.472 | 705 | 737 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 9.2E-4 | 705 | 734 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 5.84E-8 | 879 | 931 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.076 | 880 | 902 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 7.766 | 881 | 910 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0018 | 882 | 901 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.017 | 903 | 925 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.304 | 904 | 928 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 4.7E-4 | 906 | 925 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1057 aa Download sequence Send to blast |
MDILDIQQLS VEAQHRWLRP AEICEILRNY QKFRITSEPP TRPPSGSLFL FDRKVLRYFR 60 KDGHNWRKKK DGKTVKEAHE KLKVGSIDVL HCYYAHGEDN ENFQRRSYWM LEQELMHIVF 120 VHYLEVKGNR TNIGGVRETD EFSQDLSKAS PQISSSSANN CKEPSGNTDY TSPSSSLTSL 180 YEDADTGDSH QASSTYHSYS GSPQPQVGNG LLMNNIDVSF LTGPAAKNHE GHSSINGADY 240 MPHLEKDRPM YNDSPGYIVG QQKTLGLGSW EEILEQCTMG FHTDPACTGV ALEQQNMTLN 300 GLSACDSTAE KELDSSLPTH SSWQIPFEDN SLLLPKLSVD QPPNSELPCY QDDLFNTLLH 360 QQKEKPAQSN LQAQLPNAES ECLIASNADD DIYADGNMNY SLSLRRQLFD GEEGLKKVDS 420 FSRWVSKELG EVDNLQMQSS SGISWSTVEC GNVVDDSSLS PSLSQDQLFS IIDVSPKWGY 480 ADLETEVLTT GTFLKSQQEV AKYNWSCMFG EVEGPAEILV DGVLCCHAPP HRSGQVPFYV 540 TCSNRIACSE VREFDYLLGS TKGLDITNIY GSDTDEMLLH VRLERLLSLG SVKPSKNIFK 600 DVTEKKDLIN KIISLKEEEE SNQKVDLTPE NDLSQYEVKE HLLKKLIKDK LYSWLFHKVT 660 EDGKGPSVLD DNGQGALHLA AALGYNWAIK PIVTSGVSIN FRDVNGWTAL HWAALYGREQ 720 TVALLVSLGA APGAVTDPSP EFPLGRTPAD LASVTGQKGI SGFLAESSLT SYLSSLTVNE 780 PKNDCAAENS GTTAVQTVSE RTATPMNYFD MPDTLSLKDS LTAVRNATQA ADRIHLMFRM 840 QSFERRQLSD LGDDEFGLSD EHALSLLATK TSKAGPSYGQ AHAAAVHIQK KYRGWKKRKE 900 FLIIRQRIVK IQAHVRGHQV RKQYRAIIWS VGILEKIILR WRRKGSGLRG FRNDAVIKDS 960 QPDPQHMPLK DDDYDFFKEG RKQTEERLQS ALTRVKSMVQ YPEGRAQYRR LLNVVDGFRE 1020 TKVDDMALNS ETADSDENLV DIDTLLHDDT FMSIAFE |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Binds calmodulin in a calcium-dependent manner (By similarity). Involved in freezing tolerance in association with CAMTA1 and CAMTA3. Contributes together with CAMTA1 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved together with CAMTA3 and CAMTA4 in the positive regulation of a general stress response (PubMed:25039701). Involved in tolerance to aluminum. Binds to the promoter of ALMT1 transporter and contributes to the positive regulation of aluminum-induced expression of ALMT1 (PubMed:25627216). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25627216, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
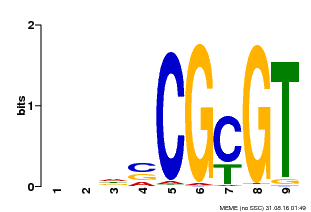
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_015886300.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt, wounding, abscisic acid, H(2)O(2) and salicylic acid (PubMed:12218065). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015886300.1 | 0.0 | calmodulin-binding transcription activator 2-like isoform X2 | ||||
| Swissprot | Q6NPP4 | 0.0 | CMTA2_ARATH; Calmodulin-binding transcription activator 2 | ||||
| TrEMBL | A0A2P5C4C4 | 0.0 | A0A2P5C4C4_PARAD; Serine/threonine protein kinase | ||||
| STRING | XP_008237378.1 | 0.0 | (Prunus mume) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



