 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_015885100.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rhamnaceae; Paliureae; Ziziphus
|
||||||||
| Family | TALE | ||||||||
| Protein Properties | Length: 382aa MW: 43558.3 Da PI: 6.6634 | ||||||||
| Description | TALE family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 29 | 1.8e-09 | 310 | 344 | 21 | 55 |
HSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHH CS
Homeobox 21 knrypsaeereeLAkklgLterqVkvWFqNrRake 55
k +yps++e+ LA+++gL+++q+ +WF N+R ++
XP_015885100.1 310 KWPYPSETEKVALAESTGLDQKQINNWFINQRKRH 344
669*****************************885 PP
| |||||||
| 2 | ELK | 37.8 | 4e-13 | 264 | 285 | 1 | 22 |
ELK 1 ELKhqLlrKYsgyLgsLkqEFs 22
ELK++LlrKYsgyL+sLkqE+s
XP_015885100.1 264 ELKNHLLRKYSGYLSSLKQELS 285
9*******************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01255 | 1.9E-21 | 120 | 164 | IPR005540 | KNOX1 |
| Pfam | PF03790 | 2.7E-22 | 121 | 162 | IPR005540 | KNOX1 |
| SMART | SM01256 | 3.3E-29 | 171 | 222 | IPR005541 | KNOX2 |
| Pfam | PF03791 | 9.0E-24 | 176 | 220 | IPR005541 | KNOX2 |
| PROSITE profile | PS51213 | 11.296 | 264 | 284 | IPR005539 | ELK domain |
| Pfam | PF03789 | 2.7E-10 | 264 | 285 | IPR005539 | ELK domain |
| SMART | SM01188 | 3.5E-7 | 264 | 285 | IPR005539 | ELK domain |
| PROSITE profile | PS50071 | 12.876 | 284 | 347 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 1.5E-19 | 285 | 359 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 1.0E-12 | 286 | 351 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 2.5E-27 | 289 | 350 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 1.43E-11 | 296 | 348 | No hit | No description |
| Pfam | PF05920 | 1.5E-16 | 304 | 343 | IPR008422 | Homeobox KN domain |
| PROSITE pattern | PS00027 | 0 | 322 | 345 | IPR017970 | Homeobox, conserved site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0001708 | Biological Process | cell fate specification | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010051 | Biological Process | xylem and phloem pattern formation | ||||
| GO:0010089 | Biological Process | xylem development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 382 aa Download sequence Send to blast |
MEEYNQMDEN TGGSRGNFLY SSAANLGGNY GRTNDGSNVG NHHHHHHQQT QMPISTFHLQ 60 SGGDCFQSSG GGGGGGHPIV KTEASTSQHL QKFHYPLMSR THQTVQHHHH ENDTTANEVE 120 AIKAKIIAHP QYCNLLEAYL DCQKVGAPPE VVARLSAARQ DFEARQRSSV SSRENSKDPE 180 LDQFMEAYYD MLVKYREELT RPIQEAMDFM RRIETQLNML SNSPVRIFGA DDKCEGIGSS 240 EDDQDNSGGE TELPEIDPRA EDRELKNHLL RKYSGYLSSL KQELSKKKKK GKLPKDARQK 300 LLSWWELHYK WPYPSETEKV ALAESTGLDQ KQINNWFINQ RKRHWKPSED MQFMVMDGLH 360 PQNAALYMDG HYMGDGHYRL GP |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probably binds to the DNA sequence 5'-TGAC-3'. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00675 | ChIP-seq | Transfer from GRMZM2G017087 | Download |
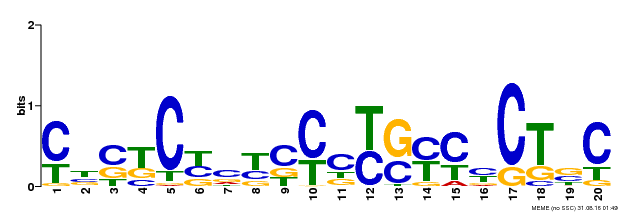
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_015885100.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015885100.1 | 0.0 | homeobox protein knotted-1-like 2 | ||||
| Swissprot | O04135 | 0.0 | KNAP2_MALDO; Homeobox protein knotted-1-like 2 | ||||
| TrEMBL | A0A061FMA1 | 0.0 | A0A061FMA1_THECC; KNOTTED-like from | ||||
| STRING | EOY15624 | 0.0 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF4584 | 32 | 46 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G08150.1 | 1e-150 | KNOTTED-like from Arabidopsis thaliana | ||||



