 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_015884823.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rhamnaceae; Paliureae; Ziziphus
|
||||||||
| Family | BBR-BPC | ||||||||
| Protein Properties | Length: 336aa MW: 37527.5 Da PI: 9.9378 | ||||||||
| Description | BBR-BPC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GAGA_bind | 399.7 | 3.8e-122 | 1 | 336 | 1 | 301 |
GAGA_bind 1 mdddgsre..rnkg.yyepa...aslkenlglqlmssiaerdakirernlalsekkaavaerdmaflqrdkalaernkalverdnkllalllven 89
md+ ++re r+kg +y+ a +++ +q+m+++aerda+i+ernla+sekkaa+aerdmaflqrd+a+aern+a++erdn++++l++++n
XP_015884823.1 1 MDEGAHREngRHKGeQYKTAqgqWLMHQPSMKQIMAIMAERDAAIQERNLAVSEKKAALAERDMAFLQRDAAIAERNNAVLERDNAIATLQYRDN 95
9999999899*******966553345556667*************************************************************** PP
GAGA_bind 90 sla....salpvgvqvlsgtksidslqq.lse.pqledsavelreeeklealpieeaaeeakekkkkkkrqrakkpkekkakkkkkksekskkkv 178
sla s++p+g+q+++g+k++++ qq +++ +++++ ++ +re++++++lpi+ a+ea ++++ k+ +ke k +++kk +k+ +kv
XP_015884823.1 96 SLAngnmSSCPPGCQISRGVKHMHHPQQhMNHmQHMNEASYSTREMHTTDSLPISPDASEAAKSRRAKR------TKEAKPISPNKKASKPPRKV 184
99999999*******************96566789999****************998887777766664......34444444444444444444 PP
GAGA_bind 179 kkesader.............................skaekksidlvlngvslDestlPvPvCsCtGalrqCYkWGnGGWqSaCCtttiSvyPL 244
k+e++d + sk+++k +dl+ln+v++Dest+P+PvCsCtG+lrqCYkWGnGGWqS+CCttt+S+yPL
XP_015884823.1 185 KRENEDLNkmtfskshewkgghdmvgagddlnkqlvvSKSDWKGQDLGLNQVAYDESTMPAPVCSCTGVLRQCYKWGNGGWQSSCCTTTMSMYPL 279
4444442224444556899**************************************************************************** PP
GAGA_bind 245 PvstkrrgaRiagrKmSqgafkklLekLaaeGydlsnpvDLkdhWAkHGtnkfvtir 301
P+++++r+aR++grKmS++af+klL++LaaeG+dlsn+vDLkdhWAkHGtn+++ti+
XP_015884823.1 280 PAVPNKRHARVGGRKMSGSAFNKLLSRLAAEGHDLSNAVDLKDHWAKHGTNRYITIK 336
********************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01226 | 7.2E-177 | 1 | 336 | IPR010409 | GAGA-binding transcriptional activator |
| Pfam | PF06217 | 1.9E-104 | 1 | 336 | IPR010409 | GAGA-binding transcriptional activator |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 336 aa Download sequence Send to blast |
MDEGAHRENG RHKGEQYKTA QGQWLMHQPS MKQIMAIMAE RDAAIQERNL AVSEKKAALA 60 ERDMAFLQRD AAIAERNNAV LERDNAIATL QYRDNSLANG NMSSCPPGCQ ISRGVKHMHH 120 PQQHMNHMQH MNEASYSTRE MHTTDSLPIS PDASEAAKSR RAKRTKEAKP ISPNKKASKP 180 PRKVKRENED LNKMTFSKSH EWKGGHDMVG AGDDLNKQLV VSKSDWKGQD LGLNQVAYDE 240 STMPAPVCSC TGVLRQCYKW GNGGWQSSCC TTTMSMYPLP AVPNKRHARV GGRKMSGSAF 300 NKLLSRLAAE GHDLSNAVDL KDHWAKHGTN RYITIK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional regulator that specifically binds to GA-rich elements (GAGA-repeats) present in regulatory sequences of genes involved in developmental processes. {ECO:0000269|PubMed:14731261}. | |||||
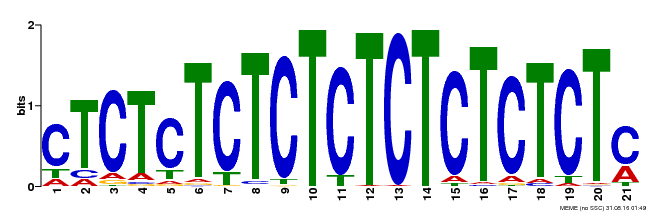
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00540 | DAP | Transfer from AT5G42520 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_015884823.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015884822.1 | 0.0 | protein BASIC PENTACYSTEINE6 | ||||
| Refseq | XP_015884824.1 | 0.0 | protein BASIC PENTACYSTEINE6 | ||||
| Refseq | XP_015884825.1 | 0.0 | protein BASIC PENTACYSTEINE6 | ||||
| Refseq | XP_015884826.1 | 0.0 | protein BASIC PENTACYSTEINE6 | ||||
| Refseq | XP_024930418.1 | 0.0 | protein BASIC PENTACYSTEINE6 | ||||
| Refseq | XP_024930420.1 | 0.0 | protein BASIC PENTACYSTEINE6 | ||||
| Refseq | XP_024930421.1 | 0.0 | protein BASIC PENTACYSTEINE6 | ||||
| Swissprot | Q8L999 | 1e-157 | BPC6_ARATH; Protein BASIC PENTACYSTEINE6 | ||||
| TrEMBL | W9RR46 | 0.0 | W9RR46_9ROSA; Uncharacterized protein | ||||
| STRING | XP_010105257.1 | 0.0 | (Morus notabilis) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G42520.1 | 1e-153 | basic pentacysteine 6 | ||||



