 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_015872669.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rhamnaceae; Paliureae; Ziziphus
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 488aa MW: 53445.7 Da PI: 7.7731 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 53.1 | 7.2e-17 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g W++eEde+l++ ++++G g+W+++++ g+ R++k+c++rw +yl
XP_015872669.1 14 KGLWSPEEDEKLLRHIAKYGHGCWSSVPKQAGLERCGKSCRLRWINYL 61
678*******************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 44.5 | 3.6e-14 | 67 | 110 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
rg ++ eE+ l++++++ lG++ W+ Ia+ ++ gRt++++k+ w++
XP_015872669.1 67 RGTFSREEENLIIELHAVLGNR-WSQIAAQLP-GRTDNDIKNLWNS 110
899*******************.*********.***********97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 1.3E-27 | 6 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 19.347 | 9 | 61 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.47E-29 | 12 | 108 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.7E-14 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.2E-15 | 14 | 61 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.42E-12 | 17 | 61 | No hit | No description |
| PROSITE profile | PS51294 | 25.992 | 62 | 116 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 3.6E-25 | 65 | 116 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 4.2E-13 | 66 | 114 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 4.0E-13 | 67 | 111 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.86E-9 | 69 | 109 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0001944 | Biological Process | vasculature development | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010089 | Biological Process | xylem development | ||||
| GO:0010119 | Biological Process | regulation of stomatal movement | ||||
| GO:0010214 | Biological Process | seed coat development | ||||
| GO:0048364 | Biological Process | root development | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 488 aa Download sequence Send to blast |
MGRHSCCYKQ KLRKGLWSPE EDEKLLRHIA KYGHGCWSSV PKQAGLERCG KSCRLRWINY 60 LRPDLKRGTF SREEENLIIE LHAVLGNRWS QIAAQLPGRT DNDIKNLWNS CLKKKLRQRG 120 VDPVTHRPIS EDKDREAKSS GEKLVSVAIS NELNLLETEN SKKAAAAAAL FGAAQVVEAN 180 NSSLCSNKTT NTTTDNNNMM MMKELFLQEQ DTSSTSNCQQ PPSDYLGQLN SLHQFKYKYG 240 STTNTGSSTT ATTRLPSSNP CSNWFLQPGK STLDMNSHDD ALSNCNSIPI ILPLSLPIPI 300 PAPSSTSFPS STASCIANCK PTSLTFPPTA TGSIDNNNSM ASFPISNNSS SSSSNCFLEN 360 NNMFGWGLVA DCSTVSEKEA HIQVNNLMDS QAGAADDIKW AEYLSNPPTF LQTTTTTQHQ 420 NSHSLLCITS SSSSTSDIKS SETQHSVMWP PPHNSKHQQL QPSQLQTTSS DICAKDIHRL 480 TAALGGPN |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 8e-29 | 12 | 116 | 5 | 108 | B-MYB |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 111 | 117 | LKKKLRQ |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00427 | DAP | Transfer from AT4G01680 | Download |
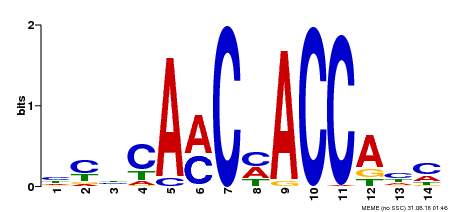
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_015872669.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015868916.1 | 0.0 | transcription factor MYB86-like | ||||
| Refseq | XP_015872487.1 | 0.0 | transcription factor MYB86-like | ||||
| Refseq | XP_015872669.1 | 0.0 | transcription factor MYB86-like | ||||
| TrEMBL | A0A2I4GGX9 | 1e-128 | A0A2I4GGX9_JUGRE; transcription factor MYB86-like | ||||
| STRING | POPTR_0014s10680.1 | 1e-119 | (Populus trichocarpa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1083 | 33 | 103 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G01680.3 | 3e-86 | myb domain protein 55 | ||||



