 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zjn_sc00068.1.g01470.1.sm.mkhc | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 685aa MW: 74537.9 Da PI: 7.324 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 75.9 | 4.7e-24 | 114 | 215 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHH CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkan 72
f k+lt+sd+++ g +++p+ +ae++ ++ + t+ +d +g +W++++iyr++++r++lt+GW+ Fv+++
Zjn_sc00068.1.g01470.1.sm.mkhc 114 FAKTLTQSDANNGGGFSVPRYCAETIfprldysADPPVQ--TVLAKDVHGMVWKFRHIYRGTPRRHLLTTGWSTFVNQK 190
89****************************996555555..8999********************************** PP
T--TT-EEEEEE-SSSEE..EEEEE-S CS
B3 73 gLkegDfvvFkldgrsefelvvkvfrk 99
+L +gD++vF+ +++++ l+v+++r+
Zjn_sc00068.1.g01470.1.sm.mkhc 191 KLVAGDSIVFM--RTENGDLCVGIRRA 215
***********..4588999****997 PP
| |||||||
| 2 | Auxin_resp | 108.8 | 5.9e-36 | 273 | 356 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrl.sGtvvgvsdldpvrWpnSk 78
aa+ a ++++FevvY+Prast+eF+vk+ v++a+++++++G Rfkmafeted s++++ +Gtv++v+ +d +rWpnS+
Zjn_sc00068.1.g01470.1.sm.mkhc 273 AANLAVSGQPFEVVYYPRASTPEFCVKAGAVRAAMRTQWCAGLRFKMAFETEDLSRISWfMGTVSAVQVADAIRWPNSP 351
688999************************************************************************* PP
Auxin_resp 79 WrsLk 83
Wr+L+
Zjn_sc00068.1.g01470.1.sm.mkhc 352 WRLLQ 356
***97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 1.7E-37 | 106 | 219 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 4.97E-29 | 109 | 216 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 2.18E-20 | 113 | 214 | No hit | No description |
| Pfam | PF02362 | 2.5E-22 | 114 | 215 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 13.084 | 114 | 216 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 1.7E-21 | 114 | 216 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 3.8E-31 | 273 | 356 | IPR010525 | Auxin response factor |
| PROSITE profile | PS51745 | 22.536 | 599 | 679 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007389 | Biological Process | pattern specification process | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0048829 | Biological Process | root cap development | ||||
| GO:0051301 | Biological Process | cell division | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 685 aa Download sequence Send to blast |
MEREREADKC LDPQLWHACA GGMVQMPPVH SKVYYFPQGH AEHAQGPVEL PAGRVPALVL 60 CRVAGVRYMA DPDTDEVFAK IRLAPVRPNE PGYADDGIGA AAAATAQEDK PASFAKTLTQ 120 SDANNGGGFS VPRYCAETIF PRLDYSADPP VQTVLAKDVH GMVWKFRHIY RGTPRRHLLT 180 TGWSTFVNQK KLVAGDSIVF MRTENGDLCV GIRRAKKGGI GGPEFLHHPP PPGGGNYGGF 240 SMFLRAEEDG NRMMASTRGK VRVRARPEEV VEAANLAVSG QPFEVVYYPR ASTPEFCVKA 300 GAVRAAMRTQ WCAGLRFKMA FETEDLSRIS WFMGTVSAVQ VADAIRWPNS PWRLLQVTWD 360 EPDLLQNVKR VNPWLVELVS NMPAIHLAPF SPPRKKLCVP LYPELPLDGH FPAPMFHGSP 420 LGRGVGPMCY FPDGTPAGIQ GARHAQFGLS LSDLHLNKLQ STLSPHGLHP LDHGAQPRIA 480 AGLIVGHPAA RDDISCLLTI GAPQNKKPAD VRKVAPPQLM LFGKPILTEQ QISLGDAGSF 540 PLAAAAKKSP SDSNAEKTVS DSDTSSPGSN QDATAENLSS GGVTLFQDTN KLLDLGLETG 600 HCKVFMQSED VGRTLDLSVF GSYEELYQRL ADMFGIEKDE LMSHVFYRDA SGTLKHTGDK 660 PFSEFTKTAR RLTILTDPSG DSLTS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldu_A | 7e-76 | 11 | 377 | 51 | 388 | Auxin response factor 5 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
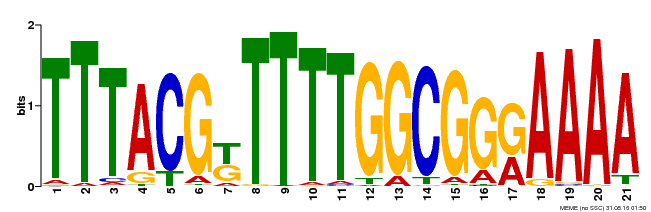
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00461 | DAP | Transfer from AT4G30080 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004966255.1 | 0.0 | auxin response factor 18 | ||||
| Refseq | XP_004966256.1 | 0.0 | auxin response factor 18 | ||||
| Swissprot | Q653H7 | 0.0 | ARFR_ORYSJ; Auxin response factor 18 | ||||
| TrEMBL | A0A3L6S3T8 | 0.0 | A0A3L6S3T8_PANMI; Auxin response factor | ||||
| STRING | Si005991m | 0.0 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP516 | 37 | 152 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G30080.1 | 0.0 | auxin response factor 16 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



