 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zjn_sc00022.1.g07910.1.am.mk | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | TCP | ||||||||
| Protein Properties | Length: 744aa MW: 79274.5 Da PI: 10.512 | ||||||||
| Description | TCP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCP | 108.6 | 9.3e-34 | 229 | 338 | 8 | 89 |
TCP 8 hskihTkvggRdRRvRlsaecaarfFdLqdeLGfdkdsktieWLlqqakpaikeltgtssssasec.....eaesssssas 83
+++h+kv+gR+RRvR+++ caar+F+L++eLG+++d++tieWLl+qa+p+i ++tg+++++a ++ +++ssss +
Zjn_sc00022.1.g07910.1.am.mk 229 GKDRHSKVNGRGRRVRMPIVCAARVFQLTRELGLKSDGQTIEWLLRQAEPSILAATGSGTTPAVFVsssapSTSSSSSHYQ 309
589*******************************************************99999444322221111111111 PP
TCP 84 nsssg..........................k 89
+ + +
Zjn_sc00022.1.g07910.1.am.mk 310 Q---SvlgkrpreegdatvaaasafwaalpsA 338
1...1234444444555555544444444442 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03634 | 6.4E-30 | 228 | 308 | IPR005333 | Transcription factor, TCP |
| PROSITE profile | PS51369 | 27.009 | 229 | 283 | IPR017887 | Transcription factor TCP subgroup |
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 744 aa Download sequence Send to blast |
MSNRSALCEP IPYRGAICGC SMTDWNDAVR TNFHQVALAR QIASKRRYSY LVSRGPRPPN 60 RACIRPRRPG DGPARDADAG LHARGQRYRA STSIQGRNLE NAISIHEVLP QQRAPLPGSF 120 SIDFDSSSTQ NPCDLSVRCP ATARQRGLER QGISALALGA PQVHTSTSRQ RYETGAMPTR 180 DVSTAAAAMF HVYQPAQIPS TAAPATAATD GTVKVPASTQ GKKKAAQGGK DRHSKVNGRG 240 RRVRMPIVCA ARVFQLTREL GLKSDGQTIE WLLRQAEPSI LAATGSGTTP AVFVSSSAPS 300 TSSSSSHYQQ SVLGKRPREE GDATVAAASA FWAALPSAAP RTEAWGFSPL EAQAATYVPM 360 AQAHHLNLLA ALSGAARRAE EETRVLDFVC ASQSVGEQFH FAPVGRFSSL SLGIASRGKG 420 TEGKRLRRLQ PAPQQSCRLP PAYENFANAN GEKVEAVASA PERTRSGFVS NPEATSSRRP 480 NCYQLTRPAM VDGCHGARPW RSGLAVACAL AAGKADPKGT WPSSEENEGG EARKRVKLPA 540 DAWTHADKTH AGLSKQCGAV LFRSPCRAPA PCCWLTTKLL VYVHRPRTSM ATHHTTPNDE 600 AESVGTRDRD ETGPCCRDFS ARDGAMPGQV RAGTPYRTAA RATSCLPTAR RVARSLSGQI 660 LFRCLASPAP TGRPRAGFGE APRRSTGRAL SSLINAVVAR QTGEEACARQ LGFGPATVLQ 720 LKSAERLSPA DCQLLGSLAL APGC |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00655 | PBM | Transfer from LOC_Os02g42380 | Download |
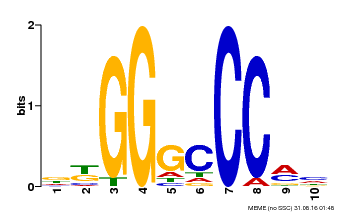
| |||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1941 | 33 | 100 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G23280.1 | 9e-30 | TCP family protein | ||||



