 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zjn_sc00003.1.g06420.1.am.mk | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | LBD | ||||||||
| Protein Properties | Length: 661aa MW: 70776.7 Da PI: 9.905 | ||||||||
| Description | LBD family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | DUF260 | 117.8 | 6.3e-37 | 384 | 476 | 1 | 92 |
DUF260 1 aCaaCkvlrrkCakdCvlapyfpaeq.pkkfanvhklFGasnvlkllkalpeeeredamsslvyeAearardPvyGavgvi 80
+C aCk+lrrkC+++C++apyf +eq +++fa+vhk+FGasnv+kll ++p ++r da+ +++yeA+ar+rdPvyG+v++i
Zjn_sc00003.1.g06420.1.am.mk 384 PCGACKFLRRKCVSGCIFAPYFDSEQgAAHFAAVHKVFGASNVSKLLLQIPAHKRLDAVVTICYEAQARLRDPVYGCVAHI 464
7***********************9989***************************************************** PP
DUF260 81 lklqqqleqlka 92
++lqqq e+++a
Zjn_sc00003.1.g06420.1.am.mk 465 FALQQQCEKKHA 476
*******98876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50891 | 20.789 | 383 | 486 | IPR004883 | Lateral organ boundaries, LOB |
| Pfam | PF03195 | 1.6E-36 | 384 | 476 | IPR004883 | Lateral organ boundaries, LOB |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010089 | Biological Process | xylem development | ||||
| GO:0010311 | Biological Process | lateral root formation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 661 aa Download sequence Send to blast |
MLLVTCRILP LRFRLRSTSS GINPKDLPAK ERFLLAWNSE LRGRVEQHLG VLHLLAVALL 60 GFQGGGSATS SLKAIEKCPG GDGPLASDLS QALASISVPD HHMDCRVHQQ LNESNVTRQH 120 PAFLEAASQR YSSSPPAGAE RSRDFVEVTR GTGKRLRVFA IRDVRQERRG GPYVAMFDRA 180 RELWHVEVKR VGERNAMRRI TWRERERKSP VLFQRNVGAR DDSGAGALAA QQWMLSDLPP 240 IGQNQTGGVQ RLFHVYTYAL PSSVLFYIAI PPGPPPFFLS GLSPLQSLLP SASIPACLPA 300 CLSEPADAAL VLSLQIERPR PASGGGSEKE QAVIRCSLLE RGRPGHVPER AMSSGGGGTS 360 TLGGGGGPSG SGGPGGGGGG GGGPCGACKF LRRKCVSGCI FAPYFDSEQG AAHFAAVHKV 420 FGASNVSKLL LQIPAHKRLD AVVTICYEAQ ARLRDPVYGC VAHIFALQQQ CEKKHASPLN 480 LESCCLPSTP GGEYYARLVR RRAIHRSRPT PPPVAPRTHH ARTHAVNLQA ELTYLQAHLA 540 TMELPSPPPP PAPPQMPVPA PFSISDLPSA SNVPSTVDLS ALFDTPPQAH QWAAVQQHHQ 600 QHQLRQPTTS YGASVRGGPG MAESSSGGGG GDLQALAREL LDRHRSGIKL ESPPPPPPHS 660 R |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5ly0_A | 4e-32 | 380 | 470 | 7 | 96 | LOB family transfactor Ramosa2.1 |
| 5ly0_B | 4e-32 | 380 | 470 | 7 | 96 | LOB family transfactor Ramosa2.1 |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
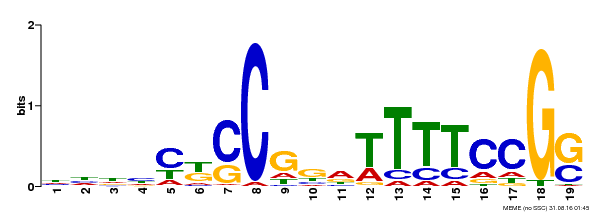
| Motif ID | Method | Source | Motif file |
| MP00314 | DAP | Transfer from AT2G45420 | Download |
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT061431 | 1e-124 | BT061431.1 Zea mays full-length cDNA clone ZM_BFb0145J09 mRNA, complete cds. | |||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3346 | 37 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G45420.1 | 5e-52 | LOB domain-containing protein 18 | ||||



