 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zjn_sc00002.1.g13170.1.am.mk | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 462aa MW: 50190.4 Da PI: 6.6221 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 55.1 | 1.3e-17 | 167 | 220 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k+++++ eq+++Le+ Fe +++ e++ +LA+ lgL+ rqV +WFqNrRa++k
Zjn_sc00002.1.g13170.1.am.mk 167 KKRRLNVEQVRTLEKNFELGNKLEPERKLQLARALGLQPRQVAIWFQNRRARWK 220
4568999**********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 123.7 | 8.7e-40 | 166 | 256 | 1 | 91 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLe 81
ekkrrl+ eqv++LE++Fe +kLeperK +lar+Lglqprqva+WFqnrRAR+ktkqlEkdy+aLkr++da+k++n++L
Zjn_sc00002.1.g13170.1.am.mk 166 EKKRRLNVEQVRTLEKNFELGNKLEPERKLQLARALGLQPRQVAIWFQNRRARWKTKQLEKDYDALKRQLDAVKADNDALV 246
69******************************************************************************* PP
HD-ZIP_I/II 82 keveeLreel 91
++++L++e
Zjn_sc00002.1.g13170.1.am.mk 247 LHNKKLQAEC 256
******9876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 3.17E-19 | 158 | 224 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.346 | 162 | 222 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.3E-17 | 165 | 226 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 5.2E-15 | 167 | 220 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 3.65E-17 | 167 | 223 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 4.3E-18 | 169 | 230 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 2.8E-5 | 193 | 202 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 197 | 220 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 2.8E-5 | 202 | 218 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 1.2E-10 | 222 | 256 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009744 | Biological Process | response to sucrose | ||||
| GO:0048826 | Biological Process | cotyledon morphogenesis | ||||
| GO:0080022 | Biological Process | primary root development | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 462 aa Download sequence Send to blast |
MGGAGGPPDS SGEGAEPCTL HVSLCVGLSL LAATPIRPMH PAPDGAVGYS FLMPMASNGM 60 ASSPPSFFPP NFLLQMQQTP LDHDPQEHHH SHLPPPPLHH PHHNPFLPSS QCPSLQDFQG 120 MAPMLGKRPM YGADVGTGDD LNGCGGGGGD DASKDELSDD GSQAGEKKRR LNVEQVRTLE 180 KNFELGNKLE PERKLQLARA LGLQPRQVAI WFQNRRARWK TKQLEKDYDA LKRQLDAVKA 240 DNDALVLHNK KLQAECSQQK GCWKRRRWEL AFHLRAYVVL TAFAESCCSK ARAASEWPAN 300 APPGLCSILP LSLDATLTQS IYSSWLLRIV LALKGGREAG SELINLNKET EASCSNRSEN 360 SSEINLDISR TPPSEGPVDP PAHQHQHAGN GAGGMILFYP SVGRSAGVDF DQLLHSSSGP 420 KMEQHGSGDG VQAPEQTSFG NLLCGVDEPP PFWPWADHQH FH |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 214 | 222 | RRARWKTKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
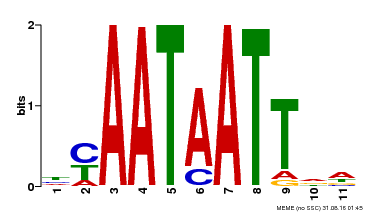
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00225 | DAP | Transfer from AT1G69780 | Download |
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC169375 | 1e-153 | AC169375.4 Sorghum bicolor clone SB_BBc0020O07, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002465782.1 | 1e-109 | homeobox-leucine zipper protein HOX21 | ||||
| Swissprot | A2XD08 | 5e-78 | HOX21_ORYSI; Homeobox-leucine zipper protein HOX21 | ||||
| TrEMBL | A3AEK5 | 1e-127 | A3AEK5_ORYSJ; Uncharacterized protein | ||||
| STRING | TRIUR3_12807-P1 | 1e-126 | (Triticum urartu) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2289 | 38 | 94 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G69780.1 | 2e-56 | HD-ZIP family protein | ||||



