 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zjn_sc00002.1.g10250.1.am.mk | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | YABBY | ||||||||
| Protein Properties | Length: 199aa MW: 22173.6 Da PI: 8.7212 | ||||||||
| Description | YABBY family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | YABBY | 54.4 | 5.3e-17 | 1 | 56 | 1 | 54 |
YABBY 1 advfssseqvCyvqCnfCntila..vsvPstslfkvvtvrCGhCtsllsvnlakas 54
+d++ se++Cyv+C+fCnt+la v vP + l+ +vtv+CGhC +l ++l+ +
Zjn_sc00002.1.g10250.1.am.mk 1 MDLVAPSEHLCYVRCTFCNTVLAlqVGVPCKRLMDTVTVKCGHCNNLSYLSLRPPT 56
578899***************9744778******************9877776655 PP
| |||||||
| 2 | YABBY | 91.5 | 2.1e-28 | 87 | 147 | 102 | 162 |
YABBY 102 senedeevprvppvirPPekrqrvPsaynrfikeeiqrikasnPdishreafsaaaknWah 162
s + e ++rvp v++PPek+ r Psaynrf++eeiqrika+ Pdi hreafs+aaknWa
Zjn_sc00002.1.g10250.1.am.mk 87 SPTSAEVSARVPFVVKPPEKKHRLPSAYNRFMREEIQRIKAAKPDIPHREAFSMAAKNWAK 147
5556778889999***********************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF04690 | 2.5E-44 | 6 | 148 | IPR006780 | YABBY protein |
| SuperFamily | SSF47095 | 1.31E-8 | 91 | 149 | IPR009071 | High mobility group box domain |
| Gene3D | G3DSA:1.10.30.10 | 1.9E-5 | 102 | 149 | IPR009071 | High mobility group box domain |
| CDD | cd01390 | 1.79E-5 | 109 | 150 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010254 | Biological Process | nectary development | ||||
| GO:0010582 | Biological Process | floral meristem determinacy | ||||
| GO:0048479 | Biological Process | style development | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 199 aa Download sequence Send to blast |
MDLVAPSEHL CYVRCTFCNT VLALQVGVPC KRLMDTVTVK CGHCNNLSYL SLRPPTVQPP 60 SPTDDPLGPF QGHCNDCRRN QPLPLGSPTS AEVSARVPFV VKPPEKKHRL PSAYNRFMRE 120 EIQRIKAAKP DIPHREAFSM AAKNWAKCDP RCSTAVSTAT STSAPEPRVV SAPQMLLQER 180 AKEQVVESFD IFKQMERSI |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Regulates carpel specification in flower development. Severe or intermediate mutation in DL causes complete or partial homeotic conversion of carpels into stamens without affecting the identities of other floral organs. Interacts antagonistically with class B genes and controls floral meristem determinacy. Regulates midrib formation in leaves probably by inducing cell proliferation in the central region of the leaf. {ECO:0000269|PubMed:14729915}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00220 | DAP | Transfer from AT1G69180 | Download |
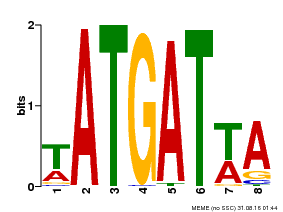
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB470268 | 1e-149 | AB470268.1 Sorghum bicolor SbDL mRNA for DL related protein, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021307124.1 | 1e-123 | protein DROOPING LEAF isoform X1 | ||||
| Swissprot | Q76EJ0 | 1e-112 | YABDL_ORYSJ; Protein DROOPING LEAF | ||||
| TrEMBL | A0A3L6TQ50 | 1e-125 | A0A3L6TQ50_PANMI; Protein DROOPING LEAF isoform X1 | ||||
| STRING | Pavir.Ia04690.1.p | 1e-124 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5951 | 36 | 57 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G69180.1 | 3e-49 | YABBY family protein | ||||



