 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zjn_sc00001.1.g05780.1.sm.mk | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | BES1 | ||||||||
| Protein Properties | Length: 313aa MW: 33490.7 Da PI: 9.5681 | ||||||||
| Description | BES1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | DUF822 | 212.2 | 1.4e-65 | 9 | 147 | 2 | 144 |
DUF822 2 gsgrkptwkErEnnkrRERrRRaiaakiyaGLRaqGnyklpkraDnneVlkALcreAGwvvedDGttyrkgskpleeaeaa 82
g gr+ptwkErEnn rRERrRRaiaaki++GLRa Gnyklpk++DnneVlkALcreAGwvvedDGttyrkg+kp+ +
Zjn_sc00001.1.g05780.1.sm.mk 9 GVGRTPTWKERENNMRRERRRRAIAAKIFTGLRALGNYKLPKHCDNNEVLKALCREAGWVVEDDGTTYRKGCKPP----PG 85
789************************************************************************....78 PP
DUF822 83 gssasaspesslq.sslkssalaspvesysaspksssfpspssldsislasaasllpvlsvls 144
+sa +sp+ss+q +s+ ss++aspv+sy+asp+sssfpsp++lds+++a ++ +lp+l+ l+
Zjn_sc00001.1.g05780.1.sm.mk 86 AASAGMSPCSSSQlLSAPSSSFASPVPSYHASPASSSFPSPTRLDSNNNA-PSLMLPFLRGLP 147
88999********9*********************************987.688888877665 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF05687 | 1.4E-61 | 10 | 134 | IPR008540 | BES1/BZR1 plant transcription factor, N-terminal |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009742 | Biological Process | brassinosteroid mediated signaling pathway | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048316 | Biological Process | seed development | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005829 | Cellular Component | cytosol | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 313 aa Download sequence Send to blast |
MTSGTPAGGV GRTPTWKERE NNMRRERRRR AIAAKIFTGL RALGNYKLPK HCDNNEVLKA 60 LCREAGWVVE DDGTTYRKGC KPPPGAASAG MSPCSSSQLL SAPSSSFASP VPSYHASPAS 120 SSFPSPTRLD SNNNAPSLML PFLRGLPTLP PLRVSNSAPV TPPLSSPTAA VSRPPTKVRR 180 LDWDAVAVVD PFRHPFFAVS APASPTRARR REHPDTIPEC DESDVSTVDS GRWISFQMAP 240 TTAPASPAYN LVNRGGGSTS NPMELNGAAP AERGRGGPEF EFDKGRVVTP WEGERIHEVA 300 AEELELTLGV GAK |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5zd4_A | 5e-27 | 12 | 85 | 372 | 445 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| 5zd4_B | 5e-27 | 12 | 85 | 372 | 445 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| 5zd4_C | 5e-27 | 12 | 85 | 372 | 445 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| 5zd4_D | 5e-27 | 12 | 85 | 372 | 445 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Positive brassinosteroid-signaling protein. Mediates downstream brassinosteroid-regulated growth response and feedback inhibition of brassinosteroid biosynthetic genes. May act as transcriptional repressor by binding the brassinosteroid-response element (5'-CGTGCG-3') in the promoter of GRAS32 (AC Q9LWU9), another positive regulator of brassinosteroid signaling (By similarity). {ECO:0000250}. | |||||
| UniProt | Positive brassinosteroid-signaling protein. Mediates downstream brassinosteroid-regulated growth response and feedback inhibition of brassinosteroid (BR) biosynthetic genes (PubMed:17699623, PubMed:19220793). May act as transcriptional repressor by binding the brassinosteroid-response element (BREE) (5'-CGTG(T/C)G-3') in the promoter of DLT (AC Q9LWU9), another positive regulator of BR signaling (PubMed:19220793). Acts as transcriptional repressor of LIC, a negative regulator of BR signaling, by binding to the BRRE element of its promoter. BZR1 and LIC play opposite roles in BR signaling and regulation of leaf bending (PubMed:22570626). {ECO:0000269|PubMed:17699623, ECO:0000269|PubMed:19220793, ECO:0000269|PubMed:22570626}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00237 | DAP | Transfer from AT1G75080 | Download |
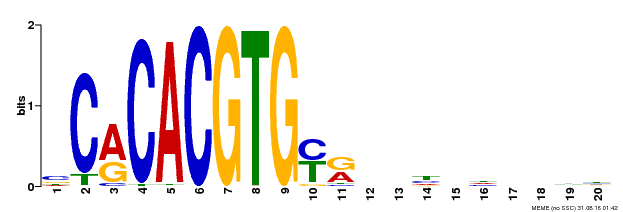
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated by 24-epibrassinolide. {ECO:0000269|PubMed:22570626}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025802391.1 | 1e-140 | protein BZR1 homolog 1-like | ||||
| Swissprot | B8B7S5 | 1e-138 | BZR1_ORYSI; Protein BZR1 homolog 1 | ||||
| Swissprot | Q7XI96 | 1e-138 | BZR1_ORYSJ; Protein BZR1 homolog 1 | ||||
| TrEMBL | A0A1E5UX51 | 1e-141 | A0A1E5UX51_9POAL; Protein BZR1-like protein 1 | ||||
| STRING | Si030508m | 1e-139 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5474 | 34 | 53 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G75080.2 | 1e-65 | BES1 family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



