 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GSVIVT01033519001 | ||||||||
| Common Name | LOC100253723 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; rosids incertae sedis; Vitales; Vitaceae; Vitis
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 350aa MW: 37831.8 Da PI: 8.9198 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 138.9 | 1.5e-43 | 51 | 126 | 1 | 76 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkk 76
+CqvegC++dls+ak y++rhkvC +hsk+p+v+v+gleqrfCqqCsrfh+l efD++krsCrrrLa+hnerrrk+
GSVIVT01033519001 51 RCQVEGCKVDLSDAKAYYSRHKVCGMHSKSPTVIVAGLEQRFCQQCSRFHQLAEFDQGKRSCRRRLAGHNERRRKP 126
6*************************************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 1.1E-35 | 45 | 113 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.436 | 49 | 126 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 3.53E-41 | 50 | 130 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 4.5E-33 | 52 | 125 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0048653 | Biological Process | anther development | ||||
| GO:2000025 | Biological Process | regulation of leaf formation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 350 aa Download sequence Send to blast |
MERGSSSLTV SSSSANSSES LNGLKFGQKI YFEDLGKVRG SGVVQGGQPP RCQVEGCKVD 60 LSDAKAYYSR HKVCGMHSKS PTVIVAGLEQ RFCQQCSRFH QLAEFDQGKR SCRRRLAGHN 120 ERRRKPPPGS LLSSRYGRLS SSIFENSSRV GGGFLMDFAA YPRHPERDTW PTTRASDRVP 180 GNQTTAMGRF LPHPWQSNSE NPLFLQGSAG GTSFHGPGIP SGECFTGASD SSCALSLLSN 240 QPWSSRNRAS GLGANSFMNP EGASMAQPTA PHSAAINHFP STSWDFKGNE GSSSSQEMPP 300 DLGLGQISQP INSQFSGGGE LPQQSGRQYM ELEHSRAYDS STQQMHWSL* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 1e-31 | 52 | 125 | 11 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Vvi.16055 | 0.0 | bud| flower| fruit| inflorescence | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Expressed constitutively during plant development, with a strong increase during flower development. {ECO:0000269|PubMed:10524240, ECO:0000269|PubMed:14573523}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
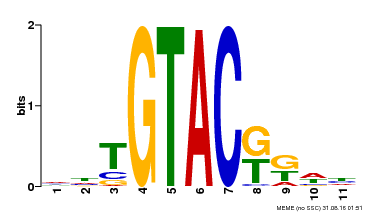
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00307 | DAP | Transfer from AT2G42200 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Negatively regulated by microRNAs miR156 and miR157. {ECO:0000305|PubMed:12202040}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HM018600 | 0.0 | HM018600.1 Vitis vinifera cultivar Xiahei promoter-binding protein SPL9 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001267898.1 | 0.0 | promoter-binding protein SPL9 | ||||
| Swissprot | Q700W2 | 1e-80 | SPL9_ARATH; Squamosa promoter-binding-like protein 9 | ||||
| TrEMBL | D6QZ29 | 0.0 | D6QZ29_VITVI; Promoter-binding protein SPL9 | ||||
| STRING | VIT_08s0007g06270.t01 | 0.0 | (Vitis vinifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP97 | 17 | 230 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G42200.1 | 1e-65 | squamosa promoter binding protein-like 9 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | GSVIVT01033519001 |
| Entrez Gene | 100253723 |



