 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GSVIVT01029458001 | ||||||||
| Common Name | VIT_09s0054g01620 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; rosids incertae sedis; Vitales; Vitaceae; Vitis
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 463aa MW: 50654.2 Da PI: 5.8246 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 102.8 | 2.1e-32 | 236 | 290 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kpr+rWtpeLHerF+eav++L G+ekAtPk +l+lm+++gLt++hvkSHLQkYRl
GSVIVT01029458001 236 KPRMRWTPELHERFLEAVNKLEGAEKATPKGVLKLMNIEGLTIYHVKSHLQKYRL 290
79****************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 9.998 | 233 | 293 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.4E-30 | 234 | 291 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 5.73E-18 | 235 | 290 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.4E-24 | 236 | 291 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 7.7E-10 | 238 | 289 | IPR001005 | SANT/Myb domain |
| Pfam | PF14379 | 3.9E-22 | 326 | 373 | IPR025756 | MYB-CC type transcription factor, LHEQLE-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 463 aa Download sequence Send to blast |
MNHHSVLSAK QTESTKGFTQ SYCAAVSPIH NLLNVELEVQ PQKLCSKSGP YSSVSSDTDA 60 QYPKCTFSRS SVFCTSLYLS SSSSTETHRP LGNLPFLPHP SMSYQSISAV HSTKTPFLSG 120 DSSGLYDEGN SEDMMKGFLN LSSDASDESF HVMNCASDNI TFSEQLELQF LSDELDIAIA 180 DNGENPRLDE IYEMPQDSST PAMALGLTVN QNHQSVAPST DASSGQPSPG AAAAHKPRMR 240 WTPELHERFL EAVNKLEGAE KATPKGVLKL MNIEGLTIYH VKSHLQKYRL AKYMPERKED 300 KKASGSEEKK AASSNNESDG RRKGNIQITE ALRLQMEVQK QLHEQLEVQR TLQLRIEEHA 360 RYLHKILEEQ QKAGSALISP PSLSSPTSPH PDSERQPSSP SATTTLPQPA ECKADSSSPP 420 PSKHKAATET TDSEQQACSK RSRLESNPEP VSDEAVVENP SQ* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 7e-26 | 236 | 293 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 7e-26 | 236 | 293 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 7e-26 | 236 | 293 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 7e-26 | 236 | 293 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 7e-26 | 236 | 293 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 7e-26 | 236 | 293 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 7e-26 | 236 | 293 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 7e-26 | 236 | 293 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Vvi.19037 | 0.0 | cell culture| inflorescence | ||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00354 | DAP | Transfer from AT3G13040 | Download |
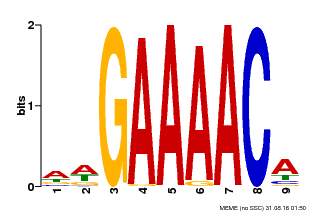
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AM482967 | 0.0 | AM482967.2 Vitis vinifera contig VV78X044419.13, whole genome shotgun sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002269813.2 | 0.0 | PREDICTED: myb family transcription factor PHL6 | ||||
| Refseq | XP_010655230.1 | 0.0 | PREDICTED: myb family transcription factor PHL6 | ||||
| Swissprot | Q949U2 | 1e-136 | PHL6_ARATH; Myb family transcription factor PHL6 | ||||
| TrEMBL | F6GYI0 | 0.0 | F6GYI0_VITVI; Uncharacterized protein | ||||
| STRING | VIT_09s0054g01620.t01 | 0.0 | (Vitis vinifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP78 | 17 | 262 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G13040.2 | 1e-105 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | GSVIVT01029458001 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



