 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GSVIVT01023149001 | ||||||||
| Common Name | LOC100264303, VIT_12s0035g01800 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; rosids incertae sedis; Vitales; Vitaceae; Vitis
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 679aa MW: 75389.1 Da PI: 6.0284 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 69.7 | 4e-22 | 128 | 228 | 1 | 98 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE- CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkld 85
f+k+lt sd++++g +++ +++a+++ +++ ++l+ d++g++W +++i+r++++r++lt+GW+ Fv++++L +gD ++F
GSVIVT01023149001 128 FCKTLTASDTSTHGGFSVLRRHADDClppldmsQQP--PWQELVAADLHGNEWHFRHIFRGQPRRHLLTTGWSVFVSSKKLVAGDAFIFL-- 215
99****************************964333..4459************************************************.. PP
SSSEE..EEEEE- CS
B3 86 grsefelvvkvfr 98
+ +++el+v+v+r
GSVIVT01023149001 216 RGENGELRVGVRR 228
449999*****97 PP
| |||||||
| 2 | Auxin_resp | 107.7 | 1.3e-35 | 254 | 337 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdld.pvrWpnSkWrsLk 83
a+ha+st++ F+v+Y+Pr+s+seF+v+++k+++a ++k+svGmRfkm+fe+e+++err+sGt+vgv d + ++ W +S+WrsLk
GSVIVT01023149001 254 ASHAISTGTLFSVFYKPRTSRSEFIVSLNKYLEARNHKLSVGMRFKMRFEGEEVPERRFSGTIVGVGDKNtSSGWADSEWRSLK 337
789***************************************************************98651678********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101936 | 5.36E-43 | 118 | 257 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 1.2E-39 | 125 | 243 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 3.20E-20 | 126 | 228 | No hit | No description |
| SMART | SM01019 | 1.6E-20 | 128 | 230 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 11.18 | 128 | 230 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 3.2E-20 | 128 | 228 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 6.9E-34 | 254 | 337 | IPR010525 | Auxin response factor |
| Pfam | PF02309 | 1.1E-10 | 550 | 643 | IPR033389 | AUX/IAA domain |
| PROSITE profile | PS51745 | 27.793 | 553 | 646 | IPR000270 | PB1 domain |
| SuperFamily | SSF54277 | 3.65E-12 | 555 | 630 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0042802 | Molecular Function | identical protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 679 aa Download sequence Send to blast |
MALVASNYPS GGPHAGAPCD ALYKELWHAC AGPLVTVPRE GERVYYFPQG HMEQLEASTT 60 HQGLDQQMPS FNLPSKILCK VVHVQLRAEP ETDEVYAQVT LLPEPDQSEI TSPDPPLPEP 120 QRCTVHSFCK TLTASDTSTH GGFSVLRRHA DDCLPPLDMS QQPPWQELVA ADLHGNEWHF 180 RHIFRGQPRR HLLTTGWSVF VSSKKLVAGD AFIFLRGENG ELRVGVRRLM RQLSNMPSSV 240 ISSHSMHLGV LATASHAIST GTLFSVFYKP RTSRSEFIVS LNKYLEARNH KLSVGMRFKM 300 RFEGEEVPER RFSGTIVGVG DKNTSSGWAD SEWRSLKVQW DEPASIFRPE RVSAWELEPL 360 VAAAAPTNLQ PAQRNKRARP PVLPSATPDL SVLGMWKSSV ESPSGFPYCD PHRGRDLYPS 420 PKFSSITKTN SFSFSGNSSP AAVSSNSMYW SNRMETATES FAPAVNKESG EKRRDTGSGC 480 RLFGFQLLDN STLEETLPVL TVGEDQPVPS LDVESDQHSE PSNINRSDIP SVSCEPDKLS 540 LRSPQESQSR QIRSCTKVHM QGIAVGRAVD LTRFDRYEDL LKKLEEMFDI QGELCGLTSI 600 WQVVYTDDED DMMMVGDDPW LEFCSMVRKI FIYTAEEVKR LSPKIKLPAM EEIKPGKLDS 660 DVAVAGTDDQ SSVVGPGC* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldv_A | 0.0 | 3 | 360 | 1 | 355 | Auxin response factor 1 |
| 4ldw_A | 0.0 | 3 | 360 | 1 | 355 | Auxin response factor 1 |
| 4ldw_B | 0.0 | 3 | 360 | 1 | 355 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Vvi.1078 | 0.0 | fruit | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Expressed in the sepals and carpels of young flower buds. At stage 10 of flower development, expression in the carpels becomes restricted to the style. Also expressed in anthers and filaments. At stage 13, expressed in the region at the top of the pedicel, including the abscission zone. {ECO:0000269|PubMed:16176952}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in the whole plant. {ECO:0000269|PubMed:10476078}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). Seems to act as transcriptional repressor. Formation of heterodimers with Aux/IAA proteins may alter their ability to modulate early auxin response genes expression. Promotes flowering, stamen development, floral organ abscission and fruit dehiscence. Acts as repressor of IAA2, IAA3 and IAA7. {ECO:0000269|PubMed:12036261, ECO:0000269|PubMed:16176952}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00097 | PBM | Transfer from AT1G59750 | Download |
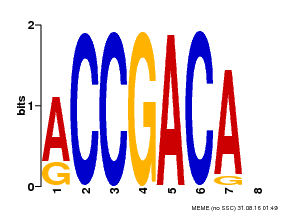
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AM476095 | 0.0 | AM476095.1 Vitis vinifera, whole genome shotgun sequence, contig VV78X038396.7, clone ENTAV 115. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010657917.1 | 0.0 | PREDICTED: auxin response factor 1 isoform X2 | ||||
| Swissprot | Q8L7G0 | 0.0 | ARFA_ARATH; Auxin response factor 1 | ||||
| TrEMBL | D7TGL8 | 0.0 | D7TGL8_VITVI; Auxin response factor | ||||
| STRING | VIT_12s0035g01800.t01 | 0.0 | (Vitis vinifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP550 | 15 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G59750.3 | 0.0 | auxin response factor 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | GSVIVT01023149001 |
| Entrez Gene | 100264303 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



