|
Plant Transcription
Factor Database
|
Transcription Factor Information
|
Basic
Information? help
Back to Top |
| TF ID |
GSVIVT01020132001 |
| Common Name | LOC100253576, VIT_01s0026g01050, VITISV_028723 |
| Organism |
|
| Taxonomic ID |
|
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; rosids incertae sedis; Vitales; Vitaceae; Vitis
|
| Family |
MYB_related |
| Protein Properties |
Length: 301aa MW: 32386.2 Da PI: 10.0734 |
| Description |
MYB_related family protein |
| Gene Model |
| Gene Model ID |
Type |
Source |
Coding Sequence |
| GSVIVT01020132001 | genome | Genoscope | View CDS |
|
| Signature Domain? help Back to Top |
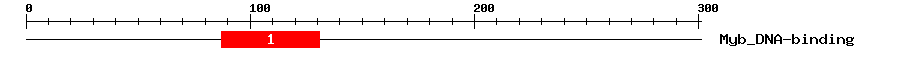 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | Myb_DNA-binding | 43.5 | 7.1e-14 | 87 | 131 | 3 | 47 |
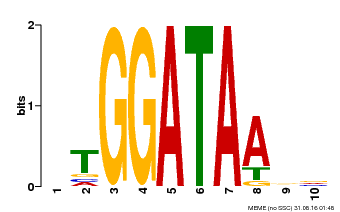
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT+eE+ l++ + ++ G+g+W+ I+r + k+Rt+ q+ s+ qky
GSVIVT01020132001 87 PWTEEEHRLFLLGLQKVGKGDWRGISRNFVKTRTPTQVASHAQKY 131
8*******************************************9 PP
|
| Gene Ontology ? help Back to Top |
| GO Term |
GO Category |
GO Description |
| GO:0006338 | Biological Process | chromatin remodeling |
| GO:0006357 | Biological Process | regulation of transcription from RNA polymerase II promoter |
| GO:0009651 | Biological Process | response to salt stress |
| GO:0009723 | Biological Process | response to ethylene |
| GO:0009733 | Biological Process | response to auxin |
| GO:0009737 | Biological Process | response to abscisic acid |
| GO:0009739 | Biological Process | response to gibberellin |
| GO:0009751 | Biological Process | response to salicylic acid |
| GO:0009753 | Biological Process | response to jasmonic acid |
| GO:0031540 | Biological Process | regulation of anthocyanin biosynthetic process |
| GO:0035066 | Biological Process | positive regulation of histone acetylation |
| GO:0046686 | Biological Process | response to cadmium ion |
| GO:0080167 | Biological Process | response to karrikin |
| GO:0005634 | Cellular Component | nucleus |
| GO:0003677 | Molecular Function | DNA binding |
| GO:0003682 | Molecular Function | chromatin binding |
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding |
| GO:0003713 | Molecular Function | transcription coactivator activity |
| GO:0004402 | Molecular Function | histone acetyltransferase activity |
| GO:0008270 | Molecular Function | zinc ion binding |
| Sequence ? help Back to Top |
| Protein Sequence Length: 301 aa
Download sequence Send
to blast |
MSRSCSQCGN NGHNSRTCTE SGGAGAAAND IMLFGVRITE GAFRKSASMT NLSQYEQPQD 60
SNADAGYASD DVVHASARSR ERKRGVPWTE EEHRLFLLGL QKVGKGDWRG ISRNFVKTRT 120
PTQVASHAQK YFLRRNNHNR RRRRSSLFDI TADTFMGSTI LEEDQVHQET VSPPQLHSHL 180
NGSAGGFPVP TFPVTMAPVV LPVASNGSME ILTLGQSNQA KGPTKLIRPI PILPVPPSSK 240
MADLNLNIPS TADPLPLSLK PSSSTSTSPQ SPTATRHSSA FQAMSESFNS STGDNIISVA 300
*
|
| Expression --
Description ? help
Back to Top |
| Source |
Description |
| Uniprot | DEVELOPMENTAL STAGE: Accumulates during leaf expansion. First observed at the tip of the leaves 12 days after sowing (DAS). At 14 DAS, expressed throughout the leaf blade to fade out thereafter in a basipetal manner. In mature leaves, detected in vascular tissue, especially in companion cells (PubMed:24806884). Accumulates to higher levels in old rosette leaves than in young rosette and cauline leaves (PubMed:25920996). {ECO:0000269|PubMed:24806884, ECO:0000269|PubMed:25920996}. |
| Uniprot | TISSUE SPECIFICITY: Expressed ubiquitously, except in hypocotyls, root tips and lateral root primordia. {ECO:0000269|PubMed:25920996}. |
| Functional Description ? help
Back to Top |
| Source |
Description |
| UniProt | Transcriptional repressor (PubMed:23888064, PubMed:24806884). Direct regulator of the transcription of peroxidase (Prxs) and reactive oxygen species (ROS)-related genes via the recognition of 5'-ATCACA-3' motif (PubMed:24806884). Binds to 5'-TATCCA-3' motif (TA box) and represses the activity of corresponding promoters (e.g. sugar response genes) (PubMed:25920996). Regulates hypocotyl elongation in response to darkness by enhancing auxin accumulation in a phytochrome-interacting factor (PIF) proteins-dependent manner. Promotes lateral roots formation (PubMed:23888064). Promotes cell expansion during leaves development via the modulation of cell wall-located Prxs (PubMed:24806884). Plays a critical role in developmentally regulated and dark-induced onset of leaf senescence by repressing the transcription of several genes involved in chloroplast function and responses to light and auxin. Promotes responses to auxin, abscisic acid (ABA), and ethylene (PubMed:25920996). {ECO:0000269|PubMed:23888064, ECO:0000269|PubMed:24806884, ECO:0000269|PubMed:25920996}. |
| Regulation -- Description ? help
Back to Top |
| Source |
Description |
| UniProt | INDUCTION: Slightly induced by CdCl(2) (PubMed:16463103). Accumulates in the dark (PubMed:23888064, PubMed:25920996). Diurnal expression pattern with maximal levels in the morning (at protein level). Specifically induced during leaf expansion (PubMed:24806884). Expressed in old and dark-treated leaves (PubMed:25920996). {ECO:0000269|PubMed:16463103, ECO:0000269|PubMed:23888064, ECO:0000269|PubMed:24806884, ECO:0000269|PubMed:25920996}. |