 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GSVIVT01019600001 | ||||||||
| Common Name | VIT_02s0025g02070 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; rosids incertae sedis; Vitales; Vitaceae; Vitis
|
||||||||
| Family | Nin-like | ||||||||
| Protein Properties | Length: 875aa MW: 96991.9 Da PI: 6.0194 | ||||||||
| Description | Nin-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | RWP-RK | 92.2 | 3.8e-29 | 559 | 609 | 2 | 52 |
RWP-RK 2 ekeisledlskyFslpikdAAkeLgvclTvLKriCRqyGIkRWPhRkiksl 52
ek+isle+l++yF++++kdAAk+Lgvc+T++KriCRq+GI+RWP+Rki+++
GSVIVT01019600001 559 EKSISLEVLQQYFAGSLKDAAKSLGVCPTTMKRICRQHGISRWPSRKINKV 609
799**********************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51519 | 17.359 | 548 | 629 | IPR003035 | RWP-RK domain |
| Pfam | PF02042 | 3.3E-26 | 562 | 609 | IPR003035 | RWP-RK domain |
| PROSITE profile | PS51745 | 22.063 | 771 | 853 | IPR000270 | PB1 domain |
| SMART | SM00666 | 8.4E-23 | 771 | 853 | IPR000270 | PB1 domain |
| Pfam | PF00564 | 2.0E-16 | 771 | 852 | IPR000270 | PB1 domain |
| SuperFamily | SSF54277 | 4.25E-22 | 771 | 855 | No hit | No description |
| Gene3D | G3DSA:3.10.20.240 | 1.1E-24 | 772 | 852 | No hit | No description |
| CDD | cd06407 | 2.54E-35 | 772 | 852 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0010118 | Biological Process | stomatal movement | ||||
| GO:0010167 | Biological Process | response to nitrate | ||||
| GO:0042128 | Biological Process | nitrate assimilation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 875 aa Download sequence Send to blast |
MSEPDEGIRK SRDCPPPSQA VDRDSFMDFD LDLDGSWPLD QISFVSNPMS PFLFSSSDQP 60 CSPLWAFSDD ADDKPSAIGV GGGLRLSECS RFLTCNPDLI PESRTENDEK RRLPPSVFTL 120 TPIENPDGCC IIKERMTQAL RYFKESTEQH VLAQVWAPVK NGDRCLLTTY GQPFVLDPHS 180 NGLHQYRMIS LTYTFSVDGE SDGALRLPAR VFRQKLPEWT PNVQYYSSRE YSRLNHALHY 240 NVRGTLALPV FEPSGPSCVG VLELIMTSQK INYAPEVDKV CKALEAVNLK SSEILEHPKA 300 QICNEGRQNA LAEILEIFTV VCETYKLPLA QTWVPCRHRS VLAGGGGLRK SCSSFDGSCM 360 GQVCMSTTDV AFYVVDAHMW GFREACAEHH LQKGQGVAGR AFESHNSCYC SNITQFCKTE 420 YPLVHYARMF GLTCCFAICL RSTHTGNDDY ILEFFLPPSI TDSRDQQTLL DSLLATMKQH 480 FQSLRVASGK EFEEEEKSVE IIKLPMNGKL DSRLESIQIS QSTPSPPGPD ILPSRGEMQQ 540 LDSTKHQLMP SERKRGKTEK SISLEVLQQY FAGSLKDAAK SLGVCPTTMK RICRQHGISR 600 WPSRKINKVN RSLSKLKRVI ESVQVSERAF GLTSLTSSPL PVAVGSISWP ATLNGPYQQN 660 SPELGKGATG SKTRSGSREE SAGTPTSHGS CQGSPENETT SAKNHSNSPI YDQSAFSIPE 720 ALITTEPQTH FGGMLIEDAG SSKDLRNLCP SVADAMLDER VPESTRPDVR TMTIKATYRD 780 DIIRFRIPLT SGIVELKEEV AKRLKLEVGT FDIKYLDDDH EWVLIACNAD LQECMDISWT 840 TGSNIIRLLV QDLMTNLGSS CESTGETNCK ILNG* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Vvi.17311 | 0.0 | root | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, stems, leaves, flowers and siliques. Detected in root hairs, emerging secondary roots, vascular tissues, leaf parenchyma cells and stomata. {ECO:0000269|PubMed:18826430}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in regulation of nitrate assimilation and in transduction of the nitrate signal. {ECO:0000269|PubMed:18826430}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00449 | DAP | Transfer from AT4G24020 | Download |
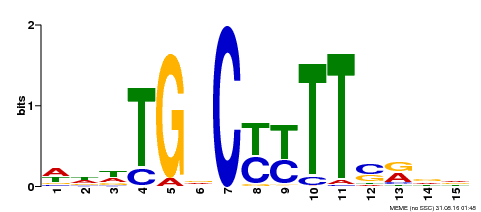
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Not regulated by the N source or by nitrate. {ECO:0000269|PubMed:18826430}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AM482982 | 0.0 | AM482982.1 Vitis vinifera, whole genome shotgun sequence, contig VV78X185475.33, clone ENTAV 115. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010659716.1 | 0.0 | PREDICTED: protein NLP6 | ||||
| Swissprot | Q84TH9 | 0.0 | NLP7_ARATH; Protein NLP7 | ||||
| TrEMBL | F6HU52 | 0.0 | F6HU52_VITVI; Uncharacterized protein | ||||
| STRING | VIT_02s0025g02070.t01 | 0.0 | (Vitis vinifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP299 | 16 | 97 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G24020.1 | 0.0 | NIN like protein 7 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | GSVIVT01019600001 |



