| Signature Domain? help Back to Top |
 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | SRF-TF | 82.1 | 3.5e-26 | 9 | 59 | 1 | 51 |
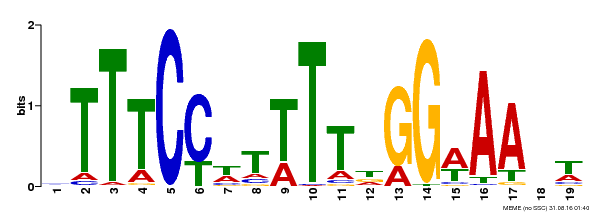
S---SHHHHHHHHHHHHHHHHHHHHHHHHHHT-EEEEEEE-TTSEEEEEE- CS
SRF-TF 1 krienksnrqvtfskRrngilKKAeELSvLCdaevaviifsstgklyeyss 51
k+ien rqvtfskRr g+lKKA+EL +LCda+v viifs+tgkl+e+ss
GSVIVT01001437001 9 KKIENANSRQVTFSKRRVGLLKKASELAILCDAQVGVIIFSNTGKLFEFSS 59
68***********************************************96 PP
|
| 2 | K-box | 75.2 | 1.8e-25 | 79 | 170 | 9 | 100 |
K-box 9 leeakaeslqqelakLkkeienLqreqRhllGedLesLslkeLqqLeqqLekslkkiRskKnellleqieelqkkekelqeenkaLrkklee 100
l e kae+ +e++ Lk+ei++Lq+ q +llG+dL+ LslkeLq+LeqqL++sl +++++K+++l+eq+e+++ +e+++ en++Lr+++ee
GSVIVT01001437001 79 LVEYKAEQEPKEVDILKDEIRKLQTRQLQLLGKDLSGLSLKELQNLEQQLNESLLSVKERKEQVLMEQLEQSRVQEQRAVLENETLRRQVEE 170
7778899999*******************************************************************************987 PP
|
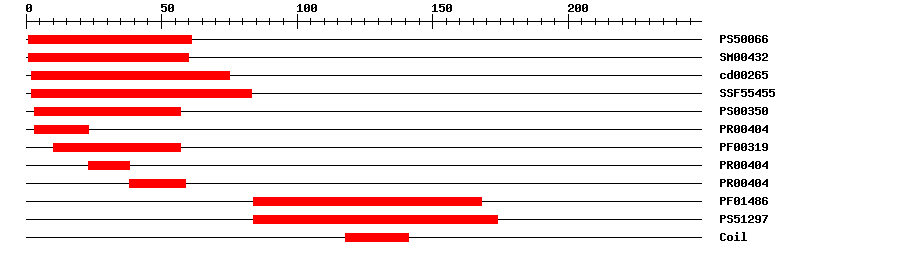
| Protein Features
? help Back to Top |
 |
| Database |
Entry ID |
E-value |
Start |
End |
InterPro ID |
Description |
| PROSITE profile | PS50066 | 31.244 | 1 | 61 | IPR002100 | Transcription factor, MADS-box |
| SMART | SM00432 | 2.9E-40 | 1 | 60 | IPR002100 | Transcription factor, MADS-box |
| CDD | cd00265 | 7.45E-41 | 2 | 75 | No hit | No description |
| SuperFamily | SSF55455 | 7.98E-32 | 2 | 83 | IPR002100 | Transcription factor, MADS-box |
| PROSITE pattern | PS00350 | 0 | 3 | 57 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 1.6E-27 | 3 | 23 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF00319 | 8.3E-25 | 10 | 57 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 1.6E-27 | 23 | 38 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 1.6E-27 | 38 | 59 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF01486 | 9.5E-20 | 84 | 168 | IPR002487 | Transcription factor, K-box |
| PROSITE profile | PS51297 | 14.418 | 84 | 174 | IPR002487 | Transcription factor, K-box |
| Expression --
Description ? help
Back to Top |
| Source |
Description |
| Uniprot | DEVELOPMENTAL STAGE: During the reproductive phase, accumulates in immature buds and at the base of the floral organs, and in the receptacle, ovules, anther filaments, and stigma and style of open flowers. Accumulates before fertilization in the cytoplasm in the cells of the egg apparatus and moves into the nucleus during early stages of development following fertilization in the suspensor, embryo, and endosperm, mainly double fertilization derived tissues (at protein level). Highly expressed in developing embryos. In young seedlings, present in the shoot and root apices, lateral root primordia and throughout the vascular system. {ECO:0000269|PubMed:10318690, ECO:0000269|PubMed:10662856, ECO:0000269|PubMed:15686521, ECO:0000269|PubMed:17521410, ECO:0000269|PubMed:8953767}. |
| Uniprot | TISSUE SPECIFICITY: Expressed at low levels in flowers and siliques. Also present in seedlings. Detected during embryogenesis and accumulates during early seed development (at protein level). Expressed in shoot apices and the base of leaf petioles. {ECO:0000269|PubMed:10318690, ECO:0000269|PubMed:10662856, ECO:0000269|PubMed:12837945, ECO:0000269|PubMed:12949148, ECO:0000269|PubMed:15686521, ECO:0000269|PubMed:16028119, ECO:0000269|PubMed:17521410, ECO:0000269|PubMed:8953767}. |
| Functional Description ? help
Back to Top |
| Source |
Description |
| UniProt | Transcription factor involved in the negative regulation of flowering, probably through the photoperiodic pathway. Acts as both an activator and a repressor of transcription. Binds DNA in a sequence-specific manner in large CArG motif 5'-CC (A/T)8 GG-3'. Participates probably in the regulation of programs active during the early stages of embryo development. Prevents premature perianth senescence and abscission, fruits development and seed desiccation. Stimulates the expression of at least DTA4, LEC2, FUS3, ABI3, AT4G38680/CSP2 and GRP2B/CSP4. Can enhance somatic embryo development in vitro. {ECO:0000269|PubMed:10318690, ECO:0000269|PubMed:10662856, ECO:0000269|PubMed:12226488, ECO:0000269|PubMed:12743119, ECO:0000269|PubMed:14615187, ECO:0000269|PubMed:15084721, ECO:0000269|PubMed:15686521, ECO:0000269|PubMed:17521410, ECO:0000269|PubMed:18305206, ECO:0000269|PubMed:19269998, ECO:0000269|PubMed:19767455, ECO:0000269|PubMed:8953767}. |