 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Vradi09g03130.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Vigna
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1080aa MW: 120418 Da PI: 5.6418 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 73.2 | 3.8e-23 | 29 | 91 | 34 | 88 |
CG-1 34 pksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvgg........vevlycyYahs 88
sgsl+L++rk++ryfrkDG++w+kkkdgktv+E+hekLK evl+cy a
Vradi09g03130.1 29 YVSGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHEKLKASMdflffslwWEVLMCYTATM 91
579***************************************999999999989998887654 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01076 | 2.3E-7 | 1 | 91 | IPR005559 | CG-1 DNA-binding domain |
| PROSITE profile | PS51437 | 28.252 | 1 | 150 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 3.0E-16 | 29 | 71 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 2.0E-4 | 491 | 579 | IPR013783 | Immunoglobulin-like fold |
| SuperFamily | SSF81296 | 1.31E-16 | 493 | 579 | IPR014756 | Immunoglobulin E-set |
| Pfam | PF01833 | 1.2E-5 | 493 | 578 | IPR002909 | IPT domain |
| SuperFamily | SSF48403 | 4.2E-16 | 677 | 786 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 2.5E-16 | 683 | 787 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.70E-12 | 692 | 781 | No hit | No description |
| SMART | SM00248 | 2800 | 693 | 722 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50297 | 17.9 | 693 | 801 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 3.2E-6 | 698 | 786 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 0.027 | 726 | 755 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 9.698 | 726 | 758 | IPR002110 | Ankyrin repeat |
| SMART | SM00015 | 0.29 | 901 | 923 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.114 | 902 | 931 | IPR000048 | IQ motif, EF-hand binding site |
| SuperFamily | SSF52540 | 3.19E-7 | 903 | 952 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| Pfam | PF00612 | 0.0022 | 904 | 922 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.017 | 924 | 946 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.413 | 925 | 949 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 4.8E-4 | 927 | 946 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1080 aa Download sequence Send to blast |
MFHITSEPPN RPPSMVSAYV TDQFITASYV SGSLFLFDRK VLRYFRKDGH NWRKKKDGKT 60 VKEAHEKLKA SMDFLFFSLW WEVLMCYTAT MPMEKKMKIF RGEAIGCLNS ECDELVYILL 120 GLYSRLDGGK QYLGTTVQVN KTNVGGKTYS GEATSDSQKG SSLSSGFPRN YGSVPSGSTD 180 SMSPTSTLTS LCEDADSEDI HQASSGSQSY HEAKCLGNDC PIDKIDARSN SSYLMHPSSG 240 DHGQFPVPGS EYIPLIQGHR AHDIASWDNA MEQSSGKDTA PSLVSSTSIP PSTSGNILEE 300 NNAVPGNLLG RKNVLIEEER VSQALHSNWQ IPFGDDTGEL PKWSLTQSLG LEFGSDYGTS 360 LLGDVTDNAG SEILAEMFTF NGELKEKSVH QNISKQYTNT PSQPATKSNS DYEVAGEASI 420 NYTLTMKRGL LDGEESLKKV DSFSRWITKE FAGVDDLHTQ SSPGISWNTD DCGDVIDDTS 480 LNLSLSQDQL FSINDFSPKW AYAESEIEVL IVGTFLKSQP MVTACNWSCM FGEVEVPAEV 540 LANGILCCQA PPHKIGRVPF YVTCSNRFAC SEVREFEYRE GFDRNIDFTE FFTSSTEMVL 600 HLRLVGLLSL SSVHTSNQVL EDDMEKRNLI FKLISLKEEE EYSSREETTV EMDATKHKLK 660 EHMFHKQVKE MLYSWLLHKV TETGKGPLVL SEEGQGVLHL VAALGYDWAI KPIITAGVNI 720 NFRDVSGWTA LHWAAYCGRE RTVAVLVSMG AATAAVTDPC PEFPEGRTAS DLASSNGHKG 780 ISGFLAESLL TSHLELLTME ESKDGRKEIS GMKAVQTVSE RTATPVLYGD IPDVICLKDS 840 LNAVRNATQA ADRIHQVYRM QSFQRKQLAK HDHDDEFGLS DQQALSLLAS RMSKSGQGEG 900 LASAAAIQIQ KKFRGWKKRK EFLIIRQRIV KIQAHVRGHQ VRKQYRTIIW SVGILEKVIL 960 RWRRKGSGLR GFRSDTVNKV IPEQPIESPK EDDYDFLKEG RKQSEARFQK ALSRVKSMVQ 1020 YPEARAQYRR VLNVVEDFRQ TKGDNLNSMN SEEAVDGVED LIDIDMLLDD ENFLPIAFD* |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
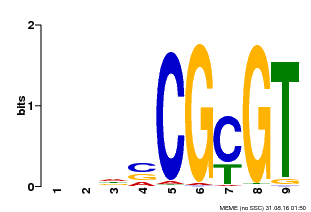
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Vradi09g03130.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015034 | 0.0 | AP015034.1 Vigna angularis var. angularis DNA, chromosome 1, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_014516371.1 | 0.0 | calmodulin-binding transcription activator 2 | ||||
| TrEMBL | A0A1S3VDD9 | 0.0 | A0A1S3VDD9_VIGRR; calmodulin-binding transcription activator 2 | ||||
| STRING | XP_007159108.1 | 0.0 | (Phaseolus vulgaris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF7406 | 26 | 42 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||



