 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Vradi09g01470.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Vigna
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 838aa MW: 91918.8 Da PI: 6.2403 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 57.7 | 2e-18 | 53 | 111 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
k ++t+eq+e+Le+l++ +++ps +r++L +++ +++ +q+kvWFqNrR +ek+
Vradi09g01470.1 53 KYVRYTPEQVEALERLYHDCPKPSSIRRQQLIRECpilsNIEPKQIKVWFQNRRCREKQ 111
5679*****************************************************97 PP
| |||||||
| 2 | START | 180.2 | 1.2e-56 | 197 | 405 | 2 | 205 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-SEEEEEEEECTT..EEEEE CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetlakaetlevissg..galql 93
+aee+++e+++ka+ ++ Wv+++ +++g++++ +++ s+++sg a+ra+g+v +++ v+e+l+d++ W ++++++++l+v+ ++ g+++l
Vradi09g01470.1 197 IAEETLAEFLSKATGTAVEWVQMPGMKPGPDSIGIVAISHGCSGVAARACGLVGLEPT-RVAEILKDRPLWFRDCRAVDVLNVLPTAngGTIEL 289
7899******************************************************.8888888888****************9999***** PP
EEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--....-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXHHHHHH CS
START 94 mvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe...sssvvRaellpSgiliepksnghskvtwvehvdlkgrlphwllrs 183
+++l+a+++l+p Rdf+ +Ry+ l +g++vi+++S+ ++q+ p+ +++vRae+lpSg+li+p+++g+s +++v+h+dl+ ++++++lr+
Vradi09g01470.1 290 LYMQLYAPTTLAPaRDFWLLRYTSVLDDGSLVICERSLKNTQNGPSmppVQHFVRAEMLPSGYLIRPCEGGGSIIHIVDHMDLEPWSVPEVLRP 383
********************************************9988899******************************************* PP
HHHHHHHHHHHHHHHHTXXXXX CS
START 184 lvksglaegaktwvatlqrqce 205
l++s+++ ++kt++a+l+++++
Vradi09g01470.1 384 LYESSTVLAQKTTMAALRHLRQ 405
*****************99876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 15.37 | 48 | 112 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.1E-15 | 50 | 116 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 1.28E-16 | 52 | 114 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 1.31E-16 | 53 | 113 | No hit | No description |
| Pfam | PF00046 | 5.8E-16 | 54 | 111 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 3.5E-18 | 55 | 111 | IPR009057 | Homeodomain-like |
| CDD | cd14686 | 2.91E-6 | 105 | 144 | No hit | No description |
| PROSITE profile | PS50848 | 25.118 | 187 | 401 | IPR002913 | START domain |
| CDD | cd08875 | 2.56E-80 | 191 | 406 | No hit | No description |
| SMART | SM00234 | 3.0E-43 | 196 | 406 | IPR002913 | START domain |
| Gene3D | G3DSA:3.30.530.20 | 2.0E-23 | 196 | 402 | IPR023393 | START-like domain |
| SuperFamily | SSF55961 | 6.04E-39 | 196 | 406 | No hit | No description |
| Pfam | PF01852 | 3.7E-54 | 197 | 405 | IPR002913 | START domain |
| Pfam | PF08670 | 1.8E-26 | 741 | 836 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 838 aa Download sequence Send to blast |
MKVVSKAKDL IPIRVSRWTE IGEAALDGAG CEIGEIDRAG FEDGSKNGID NGKYVRYTPE 60 QVEALERLYH DCPKPSSIRR QQLIRECPIL SNIEPKQIKV WFQNRRCREK QRKESSRLQA 120 VNRKLTAMNK LLMEENDRLQ KQVSQLVYEN GYFRQHTQNT TQATKDTSCE SAVTSGQHSL 180 TTQQPPRDAS PAGLLSIAEE TLAEFLSKAT GTAVEWVQMP GMKPGPDSIG IVAISHGCSG 240 VAARACGLVG LEPTRVAEIL KDRPLWFRDC RAVDVLNVLP TANGGTIELL YMQLYAPTTL 300 APARDFWLLR YTSVLDDGSL VICERSLKNT QNGPSMPPVQ HFVRAEMLPS GYLIRPCEGG 360 GSIIHIVDHM DLEPWSVPEV LRPLYESSTV LAQKTTMAAL RHLRQISHEV SQSNVTGWGR 420 RPAALRALSQ RLSRGFNEAL NGFTDEGWTT IGNDGVDDVT ILVNSSPDKL MGLNLPFANG 480 FPSVSNAVLC AKASMLLQTV ATCNFSSQNV PPAILLRFLR EHRSEWADNN MDAYTAAAIK 540 VGPCSLSGSR VGNYGGQVIL PLAHTIEHEE FLEVIKLEGV VHSPEDAIMP REMFLLQLCS 600 GMDENAVGTC AELISAPIDA SFADDAPLLP SGFRIIPLES GKEASSPNRT LDLASALDIG 660 PSGNRASNEC AGNSGCMRSV MTIAFEFAFE SHMQEHVASM ARQYVRSIIS SVQRVALALS 720 PSHLSSHTGL RSPLGTPEAQ TLAHWICNSY RCYLGVELLK SNNEGNESLL KSLWHHSDAI 780 LCCTLKALPV FTFSNQAGLD MLETTLVALQ DISLEKIFDD HGRKILFSEF PQIIQQV* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of meristem development to promote lateral organ formation. May regulates procambial and vascular tissue formation or maintenance, and vascular development in inflorescence stems. {ECO:0000269|PubMed:15598805, ECO:0000269|PubMed:15705957, ECO:0000269|PubMed:16617092}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
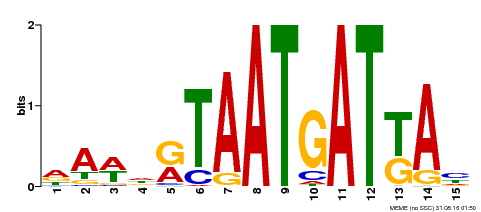
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Vradi09g01470.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. Repressed by miR165 and miR166. {ECO:0000269|PubMed:15773855, ECO:0000269|PubMed:16033795, ECO:0000269|PubMed:16617092, ECO:0000269|PubMed:17237362}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_014516466.1 | 0.0 | homeobox-leucine zipper protein ATHB-15 | ||||
| Refseq | XP_022641727.1 | 0.0 | homeobox-leucine zipper protein ATHB-15 | ||||
| Swissprot | Q9ZU11 | 0.0 | ATB15_ARATH; Homeobox-leucine zipper protein ATHB-15 | ||||
| TrEMBL | A0A1S3VDN0 | 0.0 | A0A1S3VDN0_VIGRR; homeobox-leucine zipper protein ATHB-15 | ||||
| STRING | XP_007135643.1 | 0.0 | (Phaseolus vulgaris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1946 | 33 | 87 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.3 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



