 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Vradi07g14630.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Vigna
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1517aa MW: 172491 Da PI: 9.0566 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 12.4 | 0.00048 | 1401 | 1426 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
y+C C++sF +k +L +H r+ +
Vradi07g14630.1 1401 YQCDieGCTMSFGSKQELLQHKRNiC 1426
99********************9877 PP
| |||||||
| 2 | zf-C2H2 | 12.9 | 0.00034 | 1426 | 1448 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirtH 23
Cp Cgk F ++ +L++H r+H
Vradi07g14630.1 1426 CPvkGCGKKFFSHKYLVQHRRVH 1448
9999*****************99 PP
| |||||||
| 3 | zf-C2H2 | 11.8 | 0.00074 | 1484 | 1510 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+C Cg++F+ s++ rH r+ H
Vradi07g14630.1 1484 YVCAepGCGQTFRFVSDFSRHKRKtgH 1510
899999****************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 4.0E-17 | 16 | 57 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 14.063 | 17 | 58 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 4.0E-14 | 18 | 51 | IPR003349 | JmjN domain |
| SuperFamily | SSF51197 | 7.42E-26 | 96 | 338 | No hit | No description |
| SMART | SM00558 | 5.4E-49 | 166 | 335 | IPR003347 | JmjC domain |
| PROSITE profile | PS51184 | 34.489 | 166 | 335 | IPR003347 | JmjC domain |
| Pfam | PF02373 | 1.5E-36 | 199 | 318 | IPR003347 | JmjC domain |
| Gene3D | G3DSA:3.30.160.60 | 5.7E-5 | 1388 | 1423 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 6.6 | 1401 | 1423 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 7.59E-6 | 1424 | 1460 | No hit | No description |
| PROSITE profile | PS50157 | 12.508 | 1424 | 1453 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0045 | 1424 | 1448 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1426 | 1448 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.5E-5 | 1426 | 1452 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| Gene3D | G3DSA:3.30.160.60 | 2.6E-8 | 1453 | 1478 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0016 | 1454 | 1478 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.824 | 1454 | 1483 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1456 | 1478 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.73E-9 | 1464 | 1508 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.4E-9 | 1479 | 1507 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.845 | 1484 | 1515 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.62 | 1484 | 1510 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1486 | 1510 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1517 aa Download sequence Send to blast |
MTASEEVLPW LKSLPVAPEY RPTEAEFQNP IAYIFKIEKE ASQYGICKII PPLPPSPKKT 60 ATANLTRSHP TFTTRQQQIG FCPRRPQPVR RRVWYSGHHY SLREFEAKAK AFHKTYLKKS 120 KPKPSPLELE TLYWKATLDK SFSVEYANDI PGSAFSPLRA LRDADYVGDT PWNMRVVSRG 180 SDSLLRFMKE EIPGVTSPMV YVAMLFSWFA WHVEDHDLHS LNYLHFGAPK TWYGVPKDAA 240 VAFEEVVRVH GYGGEINPLV TFAILGEKTT VMSPEVFVGA GVPCCRLVQN AGEFVVTFPR 300 AYHTGFSHGF NCGEAANIAT PEWLRFAKDA AIRRASINYP PMVSHFQLLY DLGLALCSRI 360 PGGIRAEPRS SRLKYKRNGE GETVIKELFV QDVVENNDLL HTLSKGSAIV LLPRSSSDFS 420 VCSKLRVGSR QLKVNPDFSL NVYNYEGMDS SDFISNDLMF NRNHGIKQVK SFYSVKEKFV 480 TLCERNRVLP LSSNGNIYTS SSKTDSKKET DKGDGLSDHR LFSCVTCGIL SFSCVAIVQP 540 REPAATYLMS ADCSFFNDWI VGSGVTSNKF SIAHEDASIP KPRTYTGSDN LFHNAGWTKQ 600 NAQHDSNGVP IQSVEHHAQI ADQNFEEALN SGREKGNTAL ALLASAYGNS SDSEEDQGGL 660 DIALDGDELN AVNHSASNGS QEMSSMPSHF QDPHTSPMVR VIRLNKGDDI HSRRIDNYEY 720 YMHKRLEQIM TPLNYSVKSE DHDNTSGVAF RNTRAVPHPT LNCSQDTHTD EDSSRMHIFY 780 YPKIEAEAKF VAEELGIGYT WKNTVYRQAN REDEERIQSA LDSEEAIPGN GDWAVKLGIN 840 LFYSANLSRS ALYSKQIPYN SVIYKAFGQN SPASSPTEPK VYQRRTNKQK KVVAGKWCGK 900 VWMSNQVHPL LAKRDSEDVE DETSLHGWPL PDEKIQRSES NHKSNTSTRK SGKKWKKSVE 960 KGGIWEESFV ERDSLSDNSI EDKFNKYQRR IIGGKRSRHI ERDDTASEGD YSPLPLHRKP 1020 ITKHSESSEN DGMSDDLLDD DSCIQHRRRA NTNEAKLIDS DVFSDDTMDY GSDWLRRGEL 1080 SNEQDAISDD SLSAGSLELH GKIPKGNYDK YITEEDVISD DQREVCLWKQ RGKISKGRQR 1140 SLSAKSGDGL KHHRQKQQQR NLRSRQDKHF AVENITSDDQ MEGRSFKCQR RIPKNKQVKF 1200 IEVEDVMSDD QLKGDFQKPQ RSTRRSRQNK YNDKDVMNDL AENNFHILHR TRKRKQVKGM 1260 DEDNIDSDDQ MDDILHQRHK RTLQSKQSKA EILQQTKQTN PHHARNKTSH PAKQKQGAHT 1320 KMKSRAARQT KNQSGNSKDL TLHVEEEEDG GPRTRLRKRV LEKESEGKLK EKRTKREKEK 1380 NTTAAKVLVG HAKTKDEESE YQCDIEGCTM SFGSKQELLQ HKRNICPVKG CGKKFFSHKY 1440 LVQHRRVHED DRPLKCPWKG CKMSFKWAWA RTEHIRVHTG ARPYVCAEPG CGQTFRFVSD 1500 FSRHKRKTGH ATKKNC* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 2e-80 | 8 | 349 | 6 | 346 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 2e-80 | 8 | 349 | 6 | 346 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 1250 | 1255 | TRKRKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
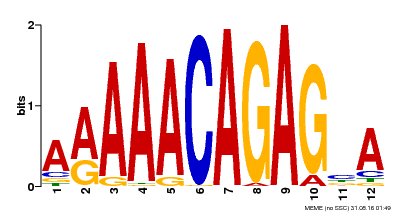
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Vradi07g14630.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015034 | 0.0 | AP015034.1 Vigna angularis var. angularis DNA, chromosome 1, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_014508868.1 | 0.0 | lysine-specific demethylase REF6-like | ||||
| Swissprot | Q9STM3 | 0.0 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | A0A1S3US05 | 0.0 | A0A1S3US05_VIGRR; lysine-specific demethylase REF6-like | ||||
| STRING | XP_007155510.1 | 0.0 | (Phaseolus vulgaris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF5648 | 32 | 50 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||



