 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Vradi06g00500.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Vigna
|
||||||||
| Family | WRKY | ||||||||
| Protein Properties | Length: 507aa MW: 55207 Da PI: 7.3715 | ||||||||
| Description | WRKY family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | WRKY | 84.5 | 1e-26 | 114 | 170 | 2 | 59 |
--SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS- CS
WRKY 2 dDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnhe 59
+DgynWrKYGqK+vkg+ef rsYY+Ct+++C++kk++++s+ ++++++ + g+Hnh+
Vradi06g00500.1 114 KDGYNWRKYGQKHVKGNEFIRSYYKCTYPNCQAKKQLQQSN-NGNITDCVSIGQHNHP 170
7***************************************9.***************8 PP
| |||||||
| 2 | WRKY | 104.8 | 4.6e-33 | 288 | 346 | 1 | 59 |
---SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS- CS
WRKY 1 ldDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnhe 59
++Dgy+WrKYGqK vkg+ +prsYYrC+ +gCpvkk+ver+++d k+v++tYeg+H+he
Vradi06g00500.1 288 VNDGYRWRKYGQKLVKGNANPRSYYRCSNPGCPVKKHVERASHDSKIVITTYEGQHDHE 346
58********************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.20.25.80 | 1.7E-23 | 107 | 172 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 19.554 | 108 | 172 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 1.96E-22 | 109 | 172 | IPR003657 | WRKY domain |
| SMART | SM00774 | 1.2E-28 | 113 | 171 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 4.5E-21 | 114 | 170 | IPR003657 | WRKY domain |
| Gene3D | G3DSA:2.20.25.80 | 1.4E-35 | 273 | 348 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 1.44E-28 | 280 | 348 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 34.475 | 283 | 348 | IPR003657 | WRKY domain |
| SMART | SM00774 | 4.7E-38 | 288 | 347 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 8.7E-26 | 289 | 345 | IPR003657 | WRKY domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009863 | Biological Process | salicylic acid mediated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 507 aa Download sequence Send to blast |
MVSSEKSEDQ NTPYDKLQHR ASPDSDVTLS QGHDTKNDLS KPEGATSIPS AVAKNEEGKD 60 SDATACALES DQEGSTCSLP PEKHLQSPDT LLNELPPLQS SQESPSIIRE KVSKDGYNWR 120 KYGQKHVKGN EFIRSYYKCT YPNCQAKKQL QQSNNGNITD CVSIGQHNHP RPQLNSTVSV 180 ECVLPVVEQA ARKPLLANVK DKTSVEQGCT SLQIKPLQSF PTAKVSPVNE LKAAHLQLSK 240 AKNQIHDNED PESKRLKKDN SNVDATGITM STREPRVVVQ TSSEVDLVND GYRWRKYGQK 300 LVKGNANPRS YYRCSNPGCP VKKHVERASH DSKIVITTYE GQHDHELPPG RTVTHNAATN 360 THTTTMNGKA GTKSEDTAVN RGEQTGLGSA SKLTEQLNGK STTKSKVGDI VEFGVSSLSN 420 EGPKIKLSEQ HQKDNSGTKD DSVSNDVICH SSSGVPCRSN EQLKGEVKPL SEGTKDCLNK 480 VAIQDTPSTE SEFNKQPAAD AEPVQS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2ayd_A | 3e-37 | 276 | 350 | 2 | 76 | WRKY transcription factor 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00094 | SELEX | Transfer from AT2G04880 | Download |
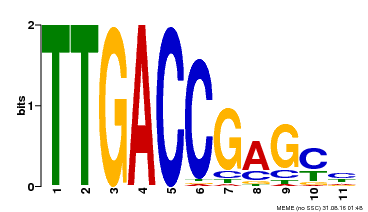
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Vradi06g00500.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015036 | 0.0 | AP015036.1 Vigna angularis var. angularis DNA, chromosome 3, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_014504627.1 | 0.0 | WRKY transcription factor 1 | ||||
| Refseq | XP_022637482.1 | 0.0 | WRKY transcription factor 1 | ||||
| TrEMBL | A0A1S3UF12 | 0.0 | A0A1S3UF12_VIGRR; WRKY transcription factor 1 | ||||
| STRING | XP_007142503.1 | 0.0 | (Phaseolus vulgaris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF9358 | 33 | 43 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G04880.2 | 2e-80 | zinc-dependent activator protein-1 | ||||



