 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Vocar.0011s0012.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; Chlorophyceae; Chlamydomonadales; Volvocaceae; Volvox
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 605aa MW: 60516.9 Da PI: 6.1315 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 56.7 | 5.6e-18 | 18 | 63 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+g+WT+eEd +l ++v G+++W++Ia++++ gR++k+c++rw++
Vocar.0011s0012.1.p 18 KGPWTPEEDTILRQLVMSQGPRNWTSIAACIP-GRSGKSCRLRWLNQ 63
79******************************.***********996 PP
| |||||||
| 2 | Myb_DNA-binding | 51.3 | 2.7e-16 | 71 | 112 | 2 | 45 |
SSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHH CS
Myb_DNA-binding 2 grWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwq 45
g++++eEd ++ a+ q+G++ W++Ia++++ gRt++++k+rw+
Vocar.0011s0012.1.p 71 GPFSAEEDAVILWAHMQYGNK-WASIAKHLP-GRTDNHIKNRWN 112
89*******************.*********.***********8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 24.601 | 13 | 68 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 3.13E-30 | 16 | 111 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 6.7E-16 | 17 | 66 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.6E-17 | 18 | 63 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 8.9E-25 | 19 | 71 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 3.28E-14 | 20 | 62 | No hit | No description |
| PROSITE profile | PS51294 | 20.019 | 69 | 119 | IPR017930 | Myb domain |
| SMART | SM00717 | 1.7E-13 | 69 | 117 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 8.2E-14 | 71 | 112 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.2E-23 | 72 | 118 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 3.53E-10 | 72 | 115 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:2000022 | Biological Process | regulation of jasmonic acid mediated signaling pathway | ||||
| GO:2000031 | Biological Process | regulation of salicylic acid mediated signaling pathway | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 605 aa Download sequence Send to blast |
MEVALFTDGA HGDDDVTKGP WTPEEDTILR QLVMSQGPRN WTSIAACIPG RSGKSCRLRW 60 LNQLSPSVKS GPFSAEEDAV ILWAHMQYGN KWASIAKHLP GRTDNHIKNR WNCTLKKKHA 120 DIMGQLRNET GPDGAVSIAA IHNVLISQQQ GGTRGPLIRG AGPTALPGKG PAAFMGPIAG 180 LGALGRDPSG APTLLPPGLP QALSGALAGA LPGGLSGLPP PLGTGLPGGL PPMMLGPMAT 240 AVAADQERRL RLAGGGGSAF SAYGNGGAKQ SDEQRGVKRS RRDGEQGTSD AESDEDSAAV 300 ALKQLAHQDP RDGNREGDAA TANNSNNNNN NNNNDGNNNN NAKGAGGAST GGTEDEDGAR 360 REEGVAPGEG AAAGREGEED NQGDRQDSEG GRGRGSDGGS GRQGAEAGPS GGSKPQRSGG 420 QPKSESTAAA GSNAANAKGG KAVPAGALPA GFTGMPSGGA GPAAGAAGDP LMQNAAAAAA 480 AAAAAAAAAS ANVPLPNSAA ANYGQLLNQL PETLMQLSST LRVVLTAGNI PAAQALVSKL 540 EELAYSVLMQ RAAFNLATAE TNTVQQHPLP QQAPVAVPAP AINGGGVDPQ ILAQLLLLSA 600 QGAS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1gv2_A | 9e-34 | 15 | 118 | 1 | 104 | MYB PROTO-ONCOGENE PROTEIN |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00596 | DAP | Transfer from AT5G67300 | Download |
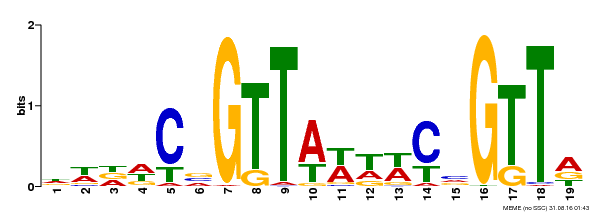
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| TrEMBL | A0A150GY24 | 1e-133 | A0A150GY24_GONPE; Uncharacterized protein | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP15 | 16 | 114 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G67300.1 | 5e-44 | myb domain protein r1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Vocar.0011s0012.1.p |



