 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Vang02g15370.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Vigna
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 283aa MW: 32221.9 Da PI: 7.2544 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 57.2 | 2.8e-18 | 86 | 138 | 4 | 56 |
-SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 4 RttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+ +++ eq+++Le+ Fe +++ e++ +LAk lgL+ rq+ +WFqNrRa++k
Vang02g15370.1 86 KKRLSLEQVKALEKSFELGNKLEPERKMQLAKALGLQPRQIAIWFQNRRARWK 138
556899**********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 128.8 | 2.2e-41 | 84 | 176 | 1 | 93 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreelke 93
ekk+rls eqvk+LE+sFe +kLeperK++la++Lglqprq+a+WFqnrRAR+ktkqlEk+ye+Lk++++a+k++n++L++++++L++el++
Vang02g15370.1 84 EKKKRLSLEQVKALEKSFELGNKLEPERKMQLAKALGLQPRQIAIWFQNRRARWKTKQLEKEYEVLKKQFEAVKADNDSLKAQNQKLHAELQT 176
69***************************************************************************************9986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.63E-19 | 76 | 142 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.151 | 80 | 140 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 4.6E-18 | 83 | 144 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 2.04E-15 | 85 | 141 | No hit | No description |
| Pfam | PF00046 | 1.7E-15 | 86 | 138 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 2.8E-19 | 87 | 147 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 9.8E-6 | 111 | 120 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 115 | 138 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 9.8E-6 | 120 | 136 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 4.1E-15 | 140 | 180 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 283 aa Download sequence Send to blast |
MAFPPSHTFM FQTHEDHDHH QLHLRSTSSS LNAFPSFPPH HFQGGGAGAG FMMKRSMSFS 60 GIENKCDEVH GDDELSDDGS QLGEKKKRLS LEQVKALEKS FELGNKLEPE RKMQLAKALG 120 LQPRQIAIWF QNRRARWKTK QLEKEYEVLK KQFEAVKADN DSLKAQNQKL HAELQTLKNR 180 DCCETGTVIS HKKDTEGSWS NGSNNSSEIN LDLSRTPVMN SPVSSQNGKS LLPTSHKPTT 240 MTQLLQSSSR SDLQDESFCN MFHNIDEQQN FWPWTEQQHQ FH* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 132 | 140 | RRARWKTKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
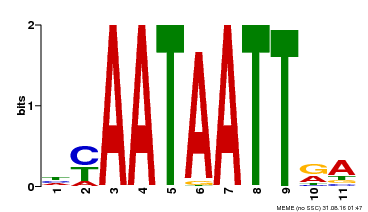
| Motif ID | Method | Source | Motif file |
| MP00322 | DAP | Transfer from AT3G01220 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Vang02g15370.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. {ECO:0000269|PubMed:12644682}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015044 | 0.0 | AP015044.1 Vigna angularis var. angularis DNA, chromosome 11, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_017437748.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein HAT7-like isoform X1 | ||||
| Swissprot | Q8LAT0 | 1e-82 | ATB20_ARATH; Homeobox-leucine zipper protein ATHB-20 | ||||
| TrEMBL | A0A0S3T7R2 | 0.0 | A0A0S3T7R2_PHAAN; Uncharacterized protein | ||||
| STRING | XP_007151217.1 | 0.0 | (Phaseolus vulgaris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1271 | 33 | 106 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G01220.1 | 1e-72 | homeobox protein 20 | ||||



