 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Vang01g08240.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Vigna
|
||||||||
| Family | GATA | ||||||||
| Protein Properties | Length: 344aa MW: 38032.7 Da PI: 7.4852 | ||||||||
| Description | GATA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GATA | 48 | 1.7e-15 | 268 | 304 | 1 | 35 |
GATA 1 CsnCgttk..TplWRrgpdgnktLCnaCGlyyrkkgl 35
C++Cg ++ Tp++Rrgp+g++tLCnaCGl+++ kg+
Vang01g08240.1 268 CRHCGISEkcTPMMRRGPEGPRTLCNACGLMWANKGT 304
**********************************997 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51320 | 14.624 | 124 | 159 | IPR010399 | Tify domain |
| SMART | SM00979 | 5.8E-11 | 124 | 159 | IPR010399 | Tify domain |
| Pfam | PF06200 | 1.6E-12 | 128 | 158 | IPR010399 | Tify domain |
| PROSITE profile | PS51017 | 13.205 | 192 | 234 | IPR010402 | CCT domain |
| Pfam | PF06203 | 7.7E-16 | 192 | 234 | IPR010402 | CCT domain |
| SMART | SM00401 | 4.3E-14 | 262 | 315 | IPR000679 | Zinc finger, GATA-type |
| PROSITE profile | PS50114 | 8.88 | 262 | 321 | IPR000679 | Zinc finger, GATA-type |
| SuperFamily | SSF57716 | 1.9E-12 | 263 | 314 | No hit | No description |
| Gene3D | G3DSA:3.30.50.10 | 6.3E-15 | 266 | 309 | IPR013088 | Zinc finger, NHR/GATA-type |
| CDD | cd00202 | 6.61E-14 | 267 | 323 | No hit | No description |
| Pfam | PF00320 | 2.2E-13 | 268 | 304 | IPR000679 | Zinc finger, GATA-type |
| PROSITE pattern | PS00344 | 0 | 268 | 295 | IPR000679 | Zinc finger, GATA-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 344 aa Download sequence Send to blast |
LCRSTHNLQF LLGFIKLYHH ITRMFALFPN LFFVLSVAKC FTMDGIHGGD SRIHISDGQH 60 PIHVPYVQEH EHHGLHHMSN GNGIDEDQND GGDTNCGGSE SVEGDIPSSH GNLPDNHAVI 120 MDQGGDAGDQ LTLSFQGQVY VFDSVSPEKV QAVLLLLGGR EIPPTMPTMP VSPHHNNRGF 180 TGTPQKFSVP QRLASLIRFR EKRKERNFDK KIRYTVRKEV ALRMQRNKGQ FTSSKSNHDE 240 SASAATNWGP NENWSAENNG SQQQDIVCRH CGISEKCTPM MRRGPEGPRT LCNACGLMWA 300 NKGTLRDLSR AAPISGPIKN ENKSLETNQI VVHRVAGEAD DLS* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that specifically binds 5'-GATA-3' or 5'-GAT-3' motifs within gene promoters. {ECO:0000250, ECO:0000269|PubMed:14966217}. | |||||
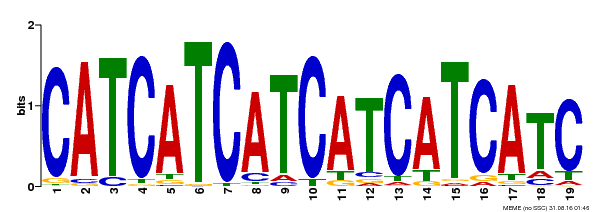
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00198 | DAP | Transfer from AT1G51600 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Vang01g08240.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015036 | 0.0 | AP015036.1 Vigna angularis var. angularis DNA, chromosome 3, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_017430361.1 | 0.0 | PREDICTED: GATA transcription factor 28 isoform X1 | ||||
| Swissprot | Q8H1G0 | 1e-134 | GAT28_ARATH; GATA transcription factor 28 | ||||
| TrEMBL | A0A4D6MHS7 | 0.0 | A0A4D6MHS7_VIGUN; Pseudo-response regulator 1 | ||||
| STRING | XP_007142122.1 | 0.0 | (Phaseolus vulgaris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF13651 | 23 | 25 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G51600.2 | 1e-137 | ZIM-LIKE 2 | ||||



