 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Vang0052ss00070.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Vigna
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1664aa MW: 187239 Da PI: 8.6814 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 12.8 | 0.00036 | 1572 | 1594 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirtH 23
Cp Cgk F ++ +L++H r+H
Vang0052ss00070.1 1572 CPvkGCGKKFFSHKYLVQHRRVH 1594
9999*****************99 PP
| |||||||
| 2 | zf-C2H2 | 11.7 | 0.00082 | 1630 | 1656 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+C Cg++F+ s++ rH r+ H
Vang0052ss00070.1 1630 YVCAepGCGQTFRFVSDFSRHKRKtgH 1656
899999****************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 2.2E-16 | 55 | 96 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 14.603 | 56 | 97 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 3.8E-14 | 57 | 90 | IPR003349 | JmjN domain |
| SuperFamily | SSF51197 | 2.66E-26 | 217 | 404 | No hit | No description |
| PROSITE profile | PS51184 | 34.037 | 217 | 386 | IPR003347 | JmjC domain |
| SMART | SM00558 | 6.9E-49 | 217 | 386 | IPR003347 | JmjC domain |
| Pfam | PF02373 | 1.6E-35 | 250 | 369 | IPR003347 | JmjC domain |
| SMART | SM00355 | 6.5 | 1547 | 1569 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.944 | 1570 | 1599 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0045 | 1570 | 1594 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 4.0E-6 | 1571 | 1598 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 4.63E-6 | 1571 | 1603 | No hit | No description |
| PROSITE pattern | PS00028 | 0 | 1572 | 1594 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.3E-8 | 1599 | 1624 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.741 | 1600 | 1629 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0014 | 1600 | 1624 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1602 | 1624 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.73E-9 | 1610 | 1654 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.5E-9 | 1625 | 1653 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.468 | 1630 | 1661 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.62 | 1630 | 1656 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1632 | 1656 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1664 aa Download sequence Send to blast |
MAEKFPSSGN RGRDANTRCE LAEASMRATE KKSGIMGGGS EGSADVLPWL KSMPVAPEYR 60 PTAAEFQDPI GYIFKIEKEA SKYGICKIIP PFPPSPKKTA IANLNRSLAV SGSTFTTRQQ 120 QIGFCPRRPR PVQRPVWQSG DHYTFTEFEF KAKSFEKAYL KRHTRKGPSP GPGLTPLETE 180 TLFWKATLDK PFSVEYANDM PGSAFSPKCR HTGDPTSLAD TPWNMRVVSR ATGSLLRFMK 240 EEIPGVTSPM VYVAMLFSWF AWHVEDHDLH SLNYLHMGAG KTWYGIPRDA AVAFEEVVRV 300 HGYGGEINPL VTFAILGEKT TVMSPEVFLS AGVPCCRLVQ NAGEFVVTFP RAYHTGFSHG 360 FNCGEAANIA TPEWLRVAKD AAIRRASLNY PPMVSHFQLL YDLALALCSR IPASISAEPR 420 SSRLKDKKKG EGETVIKELF VQDVLQNNDL LHILGKGSAV VLLPRSSVDI SVCAKLRVGS 480 QQSINVSNSE GMHSSKDFVS DDLVFNRSHG IKQEKTFYSV KDKFSMIYER NRVSSFDVNG 540 SLSTSSSKPL QRDTEGETSK EDGLSDQRLF SCVTCGILSF SCVAIVQPRD PAARYLMSAD 600 CSFFNDWVVG SGVSNSKLTT APEEATIPVP NMYTGSFSYG PIGVYWGPKG WMKKNVQDGM 660 QDVSVQSSRY ALSIESEKGN TALALLASAY GNSSDSEEDQ ISVDDHETNV LISASESLLS 720 HTQDSHASPV SALDSGDNIT LMSTSCEGLM HRRFEGNLSH QSLDHSLKKQ DYNITSGVTF 780 ENMKTVPTST SNCSQDANDA ERSLCKMSMV PFDNKNASMV LQSDEDSSRM HVFCLEHAAE 840 AEKQLRPIGG AHIFLLCHPD ILNWLRNKGT VAMKNESVVM EVISVYVQCK ESSSPNAKVK 900 FNYPKIEAEA KLVAEDLGID YTWKSIAYRH ASKEDEERIQ SALDSEEAIP GNGDWAVKLG 960 INLFYSAHLS RSPLYSKQMP YNSVIYCAFG CSSPASSPEE PKVYQRRVNR QKKVVAGKWC 1020 GKVWMSNQVH PLLAKRDSED AEDEKILLGW ILPEERIERS ESTPKGETTS RKSGKKRKST 1080 AENGRTRKVS YAKKNVVSYN STEDKPNSQP RSVHRSKKAR NVGRDRTALK GDTSAYNHRK 1140 PISKQTNFTE SDAVSDDSLD DEDDHMQHGR NFDIDNDVVS DDTGDCDSDW QQREEHSSKD 1200 DEDMERDAIS EDSEGKHDKF ISEEDIISDD QMESCFQKRQ RRTPKSRQAK YLTGKSIISD 1260 DQLELKMQKQ QRRNPKSRQA KYLNEEDIAS DDQLEGHYRR YQRRNPKGRQ AKCVAEEDEM 1320 SGDQLEDHCQ KLQTSFSRKK QNKGINREDK YVMSDDQLED HFPKQQRRIP KSRQNKYLDK 1380 EVMDDSAENN SRLLHRTPKR KQAKCMDEDD LNSDDEMEDD QKLRRTLRSK QSKPKTLQQM 1440 KQANSVHAKN QASRSIKRGS RVLVKSKTPQ QIKPRNKQSS NSREFSLLME DEEEGGPSTR 1500 LRKRATKAQG TEGKLKEKQT KRKKVKNDST GKVSVGHVKG KEEEADYQCD IDGCSMSFGS 1560 KQELLHHKKN ICPVKGCGKK FFSHKYLVQH RRVHEDERPL KCPWKGCKMT FKWAWARTEH 1620 IRVHTGARPY VCAEPGCGQT FRFVSDFSRH KRKTGHSAKK SRQ* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 2e-79 | 45 | 407 | 4 | 353 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 2e-79 | 45 | 407 | 4 | 353 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 1501 | 1523 | RKRATKAQGTEGKLKEKQTKRKK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
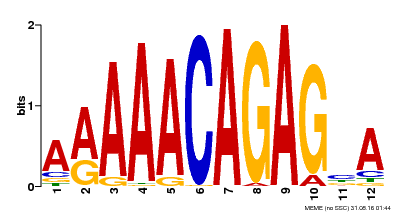
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Vang0052ss00070.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015035 | 0.0 | AP015035.1 Vigna angularis var. angularis DNA, chromosome 2, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_017421723.1 | 0.0 | PREDICTED: lysine-specific demethylase REF6-like | ||||
| Swissprot | Q9STM3 | 0.0 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | A0A0L9UCP5 | 0.0 | A0A0L9UCP5_PHAAN; Uncharacterized protein | ||||
| TrEMBL | A0A0S3RFS4 | 0.0 | A0A0S3RFS4_PHAAN; Uncharacterized protein | ||||
| STRING | XP_007137965.1 | 0.0 | (Phaseolus vulgaris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF5648 | 32 | 50 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||



