 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Vang0030ss00230.2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Vigna
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 914aa MW: 100682 Da PI: 6.4291 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 57.5 | 2.3e-18 | 92 | 150 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
k ++t+eq+e+Le+l++ +++ps +r++L +++ +++ +q+kvWFqNrR +ek+
Vang0030ss00230.2 92 KYVRYTPEQVEALERLYHDCPKPSSIRRQQLIRECpilsNIEPKQIKVWFQNRRCREKQ 150
5679*****************************************************97 PP
| |||||||
| 2 | START | 177.5 | 7.8e-56 | 236 | 444 | 2 | 205 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-SEEEEEEEECTT..EEE CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetlakaetlevissg..gal 91
+aee+++e+++ka+ ++ Wv+++ +++g++++ +++ s++++g a+ra+g+v +++ v+e+l+d++ W ++++++++l+v+ ++ g++
Vang0030ss00230.2 236 IAEETLAEFLSKATGTAVEWVQMPGMKPGPDSIGIVAISHGCTGVAARACGLVGLEPT-RVAEILKDRPSWFRDCRAVDVLNVLPTAngGTI 326
7899******************************************************.8888888888****************9999*** PP
EEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--....-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXHH CS
START 92 qlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe...sssvvRaellpSgiliepksnghskvtwvehvdlkgrlphw 179
+l +++l+a+++l+p Rdf+ +Ry+ l++g++v++++S+ ++q+ p+ +++vRae+lpSg+li+p+++g+s +++v+h+dl+ +++++
Vang0030ss00230.2 327 ELLYMQLYAPTTLAPaRDFWLLRYTSVLEDGSLVVCERSLKNTQNGPTmppVQHFVRAEMLPSGYLIRPCEGGGSIIHIVDHMDLEPWSVPE 418
**********************************************9998899*************************************** PP
HHHHHHHHHHHHHHHHHHHHTXXXXX CS
START 180 llrslvksglaegaktwvatlqrqce 205
+lr+l++s+++ ++kt++ +l+++++
Vang0030ss00230.2 419 VLRPLYESSTVLAQKTTMVALRQLRQ 444
*******************9999876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 15.402 | 87 | 151 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.2E-15 | 89 | 155 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 5.56E-17 | 89 | 155 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 1.51E-16 | 92 | 152 | No hit | No description |
| Pfam | PF00046 | 6.5E-16 | 93 | 150 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 3.9E-18 | 94 | 150 | IPR009057 | Homeodomain-like |
| CDD | cd14686 | 2.18E-6 | 144 | 183 | No hit | No description |
| PROSITE profile | PS50848 | 25.192 | 226 | 441 | IPR002913 | START domain |
| CDD | cd08875 | 3.91E-79 | 230 | 445 | No hit | No description |
| Gene3D | G3DSA:3.30.530.20 | 4.9E-23 | 233 | 441 | IPR023393 | START-like domain |
| SMART | SM00234 | 1.8E-41 | 235 | 445 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 2.61E-37 | 235 | 424 | No hit | No description |
| Pfam | PF01852 | 2.9E-53 | 236 | 444 | IPR002913 | START domain |
| Pfam | PF08670 | 1.5E-51 | 770 | 912 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 914 aa Download sequence Send to blast |
MICERRDLRS KAEVSEEPRE KGEFAESVCV GLPRMKIGEA ESFGFVLRLR EITQQVLGVG 60 GNQIVAVLSM RFVFEGMAMS CKDGKPGLDN GKYVRYTPEQ VEALERLYHD CPKPSSIRRQ 120 QLIRECPILS NIEPKQIKVW FQNRRCREKQ RKEASRLQAV NRKLTAMNKL LMEENDRLQK 180 QVSHLVYENG YFRQHTQNTA LPTKDTSCES AVTSGQRSLT AQHPPRDASP AGLMSIAEET 240 LAEFLSKATG TAVEWVQMPG MKPGPDSIGI VAISHGCTGV AARACGLVGL EPTRVAEILK 300 DRPSWFRDCR AVDVLNVLPT ANGGTIELLY MQLYAPTTLA PARDFWLLRY TSVLEDGSLV 360 VCERSLKNTQ NGPTMPPVQH FVRAEMLPSG YLIRPCEGGG SIIHIVDHMD LEPWSVPEVL 420 RPLYESSTVL AQKTTMVALR QLRQISHEVS QSNVTGWGRR PAALRALGQR LSRGFNEAIN 480 GFTDEGWSMI GNDGVDDVTV LVNSSPDKLM GLNLSFANGF PSISNAVLCA KASMLLQNVP 540 PAILLRFLRE HRSEWADNNM DAYSAAAVKV GPCGLPGTRV GNYGGQVILP LAHTIEHEEF 600 LEVIKLEGIA HSPEDAIMPR EVFLLQLCSG MDENAVGTCA ELIFAPIDAS FADDAPLLPS 660 GFRIIPLDSA KEASNPNRTL DLASALDIGP TGNRASNDYS GNSGSMRSVM TIAFEFAFES 720 HMQDHVASMA RQSVRSIISS VQRVALALSP SHLSSQAGLR TPLGTPEAQT LARWICNSYR 780 CYLGVELLKS NNEGNESLLK SLWHHSDAIL CCTLKALPVF TFANQAGLDM LETTLVALQD 840 ITLEKIFDDH GRKILCSEFP QIIQQGFACL QGGICLSSMG RPISYERVVA WKVLNEEENA 900 HCICFMFMNW SFV* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of meristem development to promote lateral organ formation. May regulates procambial and vascular tissue formation or maintenance, and vascular development in inflorescence stems. {ECO:0000269|PubMed:15598805, ECO:0000269|PubMed:15705957, ECO:0000269|PubMed:16617092}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
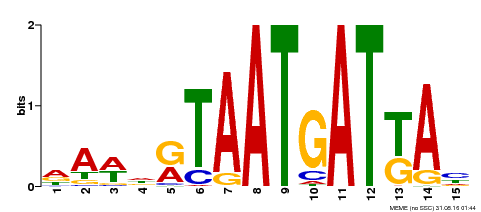
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Vang0030ss00230.2 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. Repressed by miR165 and miR166. {ECO:0000269|PubMed:15773855, ECO:0000269|PubMed:16033795, ECO:0000269|PubMed:16617092, ECO:0000269|PubMed:17237362}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_017425270.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ATHB-15 | ||||
| Swissprot | Q9ZU11 | 0.0 | ATB15_ARATH; Homeobox-leucine zipper protein ATHB-15 | ||||
| TrEMBL | A0A4D6N5B1 | 0.0 | A0A4D6N5B1_VIGUN; Homeobox-leucine zipper protein | ||||
| STRING | XP_007150614.1 | 0.0 | (Phaseolus vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



