 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 678453069 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Lentibulariaceae; Utricularia
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 843aa MW: 92766.7 Da PI: 6.0539 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 58.5 | 1.1e-18 | 52 | 110 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
k ++t+eq+e+Le+l++ +++ps +r++L +++ +++ +q+kvWFqNrR +ek+
678453069 52 KYVRYTPEQVEALERLYHDCPKPSSLRRQQLIRECpilsNIEPKQIKVWFQNRRCREKQ 110
5678*****************************************************97 PP
| |||||||
| 2 | START | 135.1 | 7.5e-43 | 196 | 373 | 2 | 173 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-SEEEEEEEECTT..EEEEEEEEXXT CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetlakaetlevissg..galqlmvaelq 99
+aee+++e+++ka+ ++ Wv+++ +++g++++ +++ s++++g a+ra+ + ++a++ e+l d++ W ++++ +++++v+s+g g+++l +++l+
678453069 196 IAEETLTEFLSKATGTAVEWVQMPGMKPGPDSIGIIAISHGCNGVASRACSLIGLEPARVAEILRDRS-IWHRDCRVVDVMNVLSTGtgGTIELLYMQLY 294
789*******************************************************8888888888.******************************* PP
TXX-SSX.EEEEEEEEEEE.TTS-E.EEEEEEEE-TTS--....-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE-- CS
START 100 alsplvp.RdfvfvRyirqlgagdw.vivdvSvdseqkppe...sssvvRaellpSgiliepksnghskvtwvehvdlk 173
al++l+p R+f+ +Ry+ + +g++ v++++S++++++ p+ ++vRae+lpSg+li+p+++g+s +++v h+dl+
678453069 295 ALTTLAPaREFWLLRYTSLMDDGSLvVVCERSLNNTANGPSiphVENFVRAEMLPSGYLIRPCDGGGSILHIVNHMDLQ 373
*******************666665166789*********9998899******************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 15.435 | 47 | 111 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 7.3E-16 | 49 | 115 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 7.27E-17 | 51 | 115 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 1.79E-16 | 52 | 112 | No hit | No description |
| Pfam | PF00046 | 3.2E-16 | 53 | 110 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 2.1E-18 | 54 | 110 | IPR009057 | Homeodomain-like |
| CDD | cd14686 | 1.64E-6 | 104 | 143 | No hit | No description |
| PROSITE profile | PS50848 | 23.158 | 186 | 392 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 8.38E-32 | 187 | 375 | No hit | No description |
| SMART | SM00234 | 8.0E-26 | 195 | 413 | IPR002913 | START domain |
| Gene3D | G3DSA:3.30.530.20 | 2.5E-16 | 195 | 374 | IPR023393 | START-like domain |
| Pfam | PF01852 | 5.6E-40 | 196 | 373 | IPR002913 | START domain |
| Pfam | PF08670 | 4.9E-44 | 701 | 842 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 843 aa Download sequence Send to blast |
MDLLIKETES EGLRKTPSEY LTSSEPNALS VVMVAVTSAY KDGGKGGVDS GKYVRYTPEQ 60 VEALERLYHD CPKPSSLRRQ QLIRECPILS NIEPKQIKVW FQNRRCREKQ RKEASRLQAV 120 NRKLTAMNKL LMEENDRLQK QISQLVYENG YFRQRTQTAT LATTDNSCDS VVTSGQHRQT 180 PQNTPQDASP SGLLSIAEET LTEFLSKATG TAVEWVQMPG MKPGPDSIGI IAISHGCNGV 240 ASRACSLIGL EPARVAEILR DRSIWHRDCR VVDVMNVLST GTGGTIELLY MQLYALTTLA 300 PAREFWLLRY TSLMDDGSLV VVCERSLNNT ANGPSIPHVE NFVRAEMLPS GYLIRPCDGG 360 GSILHIVNHM DLQISQEACH PNATAWGRRP VVLRAIGQRM SRSFNEAVNS FPDQGWSSIQ 420 GDGVDDVSLL VNSSPGKLMG SVQNYSKGLP AMSNGVLCAK ASMLLQDVSP PVLLRFLHEH 480 RSEWADTGIE ACFATAGKPG AFSLSSTRQA APFGSHVILP LAPTIENDEF MEVIRLENVS 540 NINQNMTMPN DIFLLQMYNG VDENAVGACA ELIFAPIDDT FSDDTPLLPS GFRIMPLHTK 600 PDPSTPNPTL DLASTLEIGS AGHHFSKENS AAHGASESVM TIAFQFAFQF HLQDTVATMA 660 RQYIRSIISS VQRVATALSP SNSCARAGLR ASPGTLEAEA LGQWICQSYR FYLGTEMLKP 720 QKEGNGSIFK ALWQHSDAVM CCSLKASPVF MFANQAGLDM LETSLVALQD TRLDEVFNDN 780 GRKTLLAELP QIMQQGFGYL EGGFCLSSMS RLVSYEKAVA WNVLNQEEKV HCLCIMLINW 840 SFV |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of meristem development to promote lateral organ formation. May regulates procambial and vascular tissue formation or maintenance, and vascular development in inflorescence stems. {ECO:0000269|PubMed:15598805, ECO:0000269|PubMed:15705957, ECO:0000269|PubMed:16617092}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
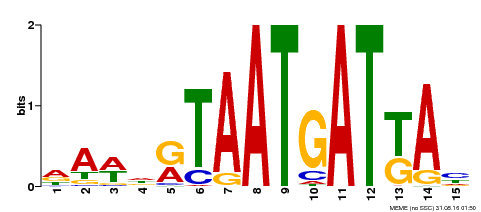
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 678453069 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. Repressed by miR165 and miR166. {ECO:0000269|PubMed:15773855, ECO:0000269|PubMed:16033795, ECO:0000269|PubMed:16617092, ECO:0000269|PubMed:17237362}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011081776.1 | 0.0 | homeobox-leucine zipper protein ATHB-15 | ||||
| Swissprot | Q9ZU11 | 0.0 | ATB15_ARATH; Homeobox-leucine zipper protein ATHB-15 | ||||
| TrEMBL | A0A4D9B785 | 0.0 | A0A4D9B785_SALSN; Homeobox-leucine zipper protein | ||||
| STRING | Migut.A00196.1.p | 0.0 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA457 | 24 | 140 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



