 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 678344382 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Lentibulariaceae; Utricularia
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1047aa MW: 116671 Da PI: 5.9084 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 176.5 | 3.4e-55 | 51 | 166 | 2 | 117 |
CG-1 2 lkekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrcywl 101
++ ++rwl++ ei++iL n++k+++++e++++p sgs++L++rk +ryfrkDG++w+kk+dgktv+E+hekLKvg+v++l+cyYah+e+n++fqrr+ywl
678344382 51 YEAQSRWLRPAEICEILRNYQKFNISAEAPSKPISGSVFLFDRKALRYFRKDGHNWRKKRDGKTVKEAHEKLKVGSVDMLHCYYAHGEDNENFQRRSYWL 150
56699*********************************************************************************************** PP
CG-1 102 Leeelekivlvhylev 117
Le +l +iv+vhyle
678344382 151 LEPDLMHIVFVHYLEI 166
*************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 78.957 | 46 | 172 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 2.3E-78 | 50 | 167 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 1.1E-49 | 52 | 165 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 5.4E-6 | 464 | 553 | IPR013783 | Immunoglobulin-like fold |
| Pfam | PF01833 | 5.7E-5 | 466 | 551 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 1.54E-14 | 466 | 552 | IPR014756 | Immunoglobulin E-set |
| PROSITE profile | PS50297 | 17.582 | 648 | 755 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 2.33E-18 | 652 | 756 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 1.0E-17 | 654 | 757 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 5.1E-8 | 656 | 720 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 3.04E-14 | 659 | 753 | No hit | No description |
| SMART | SM00248 | 1700 | 661 | 690 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 8.549 | 661 | 693 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.0028 | 694 | 723 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 9.698 | 694 | 726 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 4100 | 733 | 762 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 1.38E-6 | 861 | 908 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.036 | 864 | 886 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 7.84 | 865 | 894 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0023 | 867 | 885 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 8.4E-4 | 887 | 909 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.102 | 888 | 913 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 4.0E-5 | 889 | 908 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1047 aa Download sequence Send to blast |
MDESGSVFEL GTVSTWFSVV GFVYWGGGGC CCDVLLCLSC LEILDIQRTL YEAQSRWLRP 60 AEICEILRNY QKFNISAEAP SKPISGSVFL FDRKALRYFR KDGHNWRKKR DGKTVKEAHE 120 KLKVGSVDML HCYYAHGEDN ENFQRRSYWL LEPDLMHIVF VHYLEIKGHK ANISGGFGDA 180 GVALNPGRDS SLSSGFRGTS PSSPLSSAYA DAESENYHQA TSELHSYSEP SVQDDGHYAH 240 PSSYSSPLPA GVYIFRTQAV YGTSSVPFPL DDENGDFGGG IYSSAAATPI DAAYWPEIGR 300 NPVAGKVIYR QESASTVPSS HVNRVGLNQT FSDVSFPPDQ LNDMRLFHEY HDEKNGNLPL 360 LTSNQGIGIT SILNFESGMG SESYTFPMKK PMLGSLHSEE ALKKVDSFTR WMAKELGDAD 420 ELAVQPSGGI SWSMMGSDYE SNIPVQLQNG VDTMSPSISQ DQLFSITFFS PHWTYSDCET 480 KVLITGMFLK NEADLSESRF SIMFGQVEVP AEVLADGILC CAAPYSSPGS VPLYVTCSNR 540 LACSEIREFE YRARVGPMSS PIEVGSVAVM HLYQRFETIL GLEPHGSPQS CTGNDAQEQI 600 MNLRIISQEE ENSLEKRITS EDDVFLLGDL LIEKKLKDRF FAWILLRVAE DGKGLTLLDK 660 FGQSALHLAA ALGFIWAFEP FIIAGVKIDF RDKYGWTALH WAAFYGREEA VAVLVSLGAA 720 PGVLTHPSSE YPLSRTPADL ASANGHKGIS GFLAEISLTT HLSTLKVNDE GTTSISDVKA 780 VETLSDRVAS RTKEDDVPDK LSLKDSLAAV CNATQAAARI HQFFRIQSFQ RKQFLERESI 840 ELLDADKQAV RSGRTSRLGQ SFGKANAAAT HIQKKYRGWK KRKEFLLLRQ KAVKIQAHFR 900 GHQARKKHNI IWSVGIMEKA ILRWRRKASG LRGFRNEVPT ESTMEGSSTT LPSEEVYEVL 960 KEGRRQTEVR MQKALDRVKS MAQYPEARAQ YRRLLTVAEG FREAKDASDA MILEALEDMN 1020 YPDYPDDDLF DIASLLEDEA FMSLTYQ |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
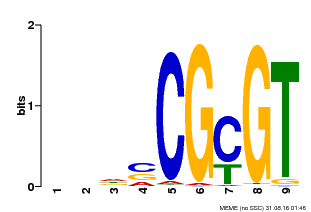
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 678344382 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011092424.1 | 0.0 | calmodulin-binding transcription activator 2 | ||||
| TrEMBL | A0A2G9H0E6 | 0.0 | A0A2G9H0E6_9LAMI; Uncharacterized protein | ||||
| STRING | Migut.B00609.1.p | 0.0 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA2230 | 21 | 46 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||



