 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 678321123 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Lentibulariaceae; Utricularia
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 260aa MW: 28461.9 Da PI: 7.9924 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 34.2 | 5.8e-11 | 11 | 55 | 3 | 45 |
SS-HHHHHHHHHHHHHTTTT...-HHHHHHHHTTTS-HHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGgg...tWktIartmgkgRtlkqcksrwq 45
+WT eEd+l+ +a+ + + +W+++a++++ g++ +++k ++
678321123 11 SWTREEDKLFERAIVTYRDEapdRWEKVASEVP-GKSVEEVKQHYE 55
7*****************99*************.**********96 PP
| |||||||
| 2 | Myb_DNA-binding | 41.6 | 2.9e-13 | 104 | 148 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT++E+ l++ + +++G+g+W+ I+r + +Rt+ q+ s+ qky
678321123 104 AWTEDEHRLFLLGLEKYGKGDWRGISRNFVVTRTPTQVASHAQKY 148
6*******************************************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51293 | 13.117 | 7 | 62 | IPR017884 | SANT domain |
| SMART | SM00717 | 8.1E-11 | 8 | 60 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.5E-6 | 10 | 58 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 1.72E-12 | 10 | 66 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 1.1E-9 | 11 | 54 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.13E-10 | 11 | 58 | No hit | No description |
| PROSITE profile | PS51294 | 17.051 | 96 | 153 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 9.33E-17 | 99 | 154 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 2.2E-10 | 101 | 151 | IPR001005 | SANT/Myb domain |
| TIGRFAMs | TIGR01557 | 1.8E-17 | 101 | 151 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 3.5E-11 | 104 | 148 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.3E-10 | 104 | 147 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 3.18E-10 | 104 | 149 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 260 aa Download sequence Send to blast |
MPAGEQSSCS SWTREEDKLF ERAIVTYRDE APDRWEKVAS EVPGKSVEEV KQHYELLIED 60 VNRIDSGFVE LPCYSVCSDG FDESGQGGTK SLSRSDQERR KGIAWTEDEH RLFLLGLEKY 120 GKGDWRGISR NFVVTRTPTQ VASHAQKYFI RLNSMNKNRR RSSIHDITSV NTGEVSVPQG 180 PITGKTNAHQ PSMGVYGTGQ CIGGPAVGTP VNLPAPGHMA YGLGAPVPGS AVGMAPMARP 240 QVVSLGHESV TRGTKSWQNK |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2cjj_A | 2e-16 | 3 | 74 | 2 | 73 | RADIALIS |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that coordinates abscisic acid (ABA) biosynthesis and signaling-related genes via binding to the specific promoter motif 5'-(A/T)AACCAT-3'. Represses ABA-mediated salt (e.g. NaCl and KCl) stress tolerance. Regulates leaf shape and promotes vegetative growth. {ECO:0000269|PubMed:26243618}. | |||||
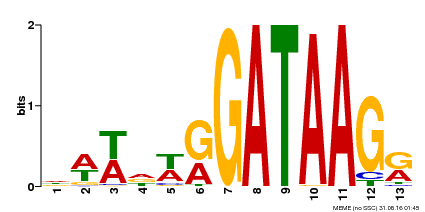
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00500 | DAP | Transfer from AT5G08520 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 678321123 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by salicylic acid (SA) and gibberellic acid (GA) (PubMed:16463103). Triggered by dehydration and salt stress (PubMed:26243618). {ECO:0000269|PubMed:16463103, ECO:0000269|PubMed:26243618}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011070471.1 | 1e-117 | transcription factor SRM1 | ||||
| Refseq | XP_011070472.1 | 1e-117 | transcription factor SRM1 | ||||
| Refseq | XP_020547442.1 | 1e-117 | transcription factor SRM1 | ||||
| Refseq | XP_020547443.1 | 1e-117 | transcription factor SRM1 | ||||
| Swissprot | Q9FNN6 | 1e-100 | SRM1_ARATH; Transcription factor SRM1 | ||||
| TrEMBL | A0A067G2J5 | 1e-112 | A0A067G2J5_CITSI; Uncharacterized protein | ||||
| STRING | XP_006484484.1 | 1e-112 | (Citrus sinensis) | ||||
| STRING | XP_006437648.1 | 1e-112 | (Citrus clementina) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA4149 | 24 | 45 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G08520.1 | 1e-102 | MYB family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



