 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 678320491 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Lentibulariaceae; Utricularia
|
||||||||
| Family | TALE | ||||||||
| Protein Properties | Length: 402aa MW: 45279.5 Da PI: 6.7249 | ||||||||
| Description | TALE family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 28 | 3.7e-09 | 328 | 363 | 20 | 55 |
HHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHH CS
Homeobox 20 eknrypsaeereeLAkklgLterqVkvWFqNrRake 55
+k +yps++e+ LA+ +gL+++q+ +WF N+R ++
678320491 328 YKWPYPSESEKVALAEATGLDQKQINNWFINQRKRH 363
4679*****************************885 PP
| |||||||
| 2 | ELK | 38.4 | 2.7e-13 | 283 | 304 | 1 | 22 |
ELK 1 ELKhqLlrKYsgyLgsLkqEFs 22
ELK++LlrKYsgyL+sL+qE+s
678320491 283 ELKNHLLRKYSGYLSSLRQELS 304
9*******************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01255 | 3.2E-20 | 114 | 158 | IPR005540 | KNOX1 |
| Pfam | PF03790 | 1.7E-21 | 115 | 156 | IPR005540 | KNOX1 |
| SMART | SM01256 | 4.8E-25 | 166 | 239 | IPR005541 | KNOX2 |
| Pfam | PF03791 | 1.2E-14 | 203 | 238 | IPR005541 | KNOX2 |
| SMART | SM01188 | 2.0E-7 | 283 | 304 | IPR005539 | ELK domain |
| PROSITE profile | PS51213 | 11.488 | 283 | 303 | IPR005539 | ELK domain |
| Pfam | PF03789 | 1.3E-10 | 283 | 304 | IPR005539 | ELK domain |
| PROSITE profile | PS50071 | 13.021 | 303 | 366 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 5.99E-20 | 304 | 380 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 3.4E-13 | 305 | 370 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 1.6E-27 | 308 | 369 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 1.27E-12 | 315 | 367 | No hit | No description |
| Pfam | PF05920 | 1.3E-16 | 323 | 362 | IPR008422 | Homeobox KN domain |
| PROSITE pattern | PS00027 | 0 | 341 | 364 | IPR017970 | Homeobox, conserved site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0001708 | Biological Process | cell fate specification | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010051 | Biological Process | xylem and phloem pattern formation | ||||
| GO:0010089 | Biological Process | xylem development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 402 aa Download sequence Send to blast |
MEGYTHPSHL TENVNNNNSD NNNYSRAGFV YTAGGGPVVA PSSSSLPFQG RVSPGSGGDH 60 LHQLEFGGLG VCFDQSQHHT VITEPGFSYN NNTDKFHYAA SSAQRQPVEL FQADAVKAKI 120 VAHPQYSNLL EAYMDCHKVG APPEVVARLA QVRQDFEAWH RGANRDSDGN SKDPELDQFM 180 VVIIGNSSSK IKLAPICLLR YQEAYYDMLV KYREELTRPL QEAMEFMRRV ETQLSMLTTA 240 PTRGFNSDDN SVGGGSSDDD QENSVGVEAG GVPGIDPRAE DRELKNHLLR KYSGYLSSLR 300 QELSKKKKKG KLPKEARQKL LSWWELHYKW PYPSESEKVA LAEATGLDQK QINNWFINQR 360 KRHWKPSEDM QFMVMDGFHH PQNPTLYMDG HYLGEGPYRL AP |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Appears to be involved in meristem formation and in the regulation of leaf morphology. Misexpression makes the leaf more compound which is always associated with growth retardation and loss of apical dominance, resulting in dwarfed, bushy plants. Probably binds to the DNA sequence 5'-TGAC-3'. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00675 | ChIP-seq | Transfer from GRMZM2G017087 | Download |
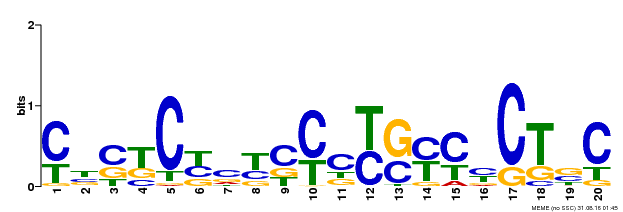
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 678320491 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011074319.1 | 1e-152 | homeotic protein knotted-1 | ||||
| Refseq | XP_022870906.1 | 1e-152 | homeotic protein knotted-1 isoform X2 | ||||
| Swissprot | Q41330 | 1e-136 | KN1_SOLLC; Homeotic protein knotted-1 | ||||
| TrEMBL | A0A2G9GBL0 | 1e-150 | A0A2G9GBL0_9LAMI; Transcription factor MEIS1 | ||||
| STRING | EOY15624 | 1e-146 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA4973 | 23 | 33 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G08150.1 | 1e-119 | KNOTTED-like from Arabidopsis thaliana | ||||



