 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | TRIUR3_32014-P1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Triticum
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 373aa MW: 40624 Da PI: 7.7277 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 36.7 | 7.6e-12 | 199 | 241 | 7 | 54 |
HHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHH CS
HLH 7 erErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksL 54
+Er+RR+++N++f +Lr ++P+ K++Ka+ L A+ YI++L
TRIUR3_32014-P1 199 TAERQRREKLNQRFYTLRAVVPNV-----SKMDKASLLGDAISYINEL 241
58*********************6.....5***************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 2.4E-9 | 15 | 64 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 16.069 | 192 | 241 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 6.94E-17 | 198 | 266 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 7.9E-14 | 198 | 247 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 3.3E-17 | 199 | 263 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 3.35E-12 | 200 | 242 | No hit | No description |
| Pfam | PF00010 | 2.4E-9 | 200 | 241 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006952 | Biological Process | defense response | ||||
| GO:0009718 | Biological Process | anthocyanin-containing compound biosynthetic process | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043425 | Molecular Function | bHLH transcription factor binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 373 aa Download sequence Send to blast |
MAMVVDGAGP AGLASAPCER ARQAYTFGLR TMVCIPLGTG VLELGATEVI FQTNDSLGRI 60 RSLFNLNGGG GGSGSWPPIA PPPQEAETDP SVLWLADAPA GDMKESPPSV EISVSKPPQP 120 QPPQIHQFEN GSTSTLTENP GLSVHAQQPP PQQAAAAAQR QNQHQLQHQH QLQLQHQHNQ 180 GPFRRELNFS DFASNASVTA ERQRREKLNQ RFYTLRAVVP NVSKMDKASL LGDAISYINE 240 LRGKMTALES DKETLHSQIE ALKKERDARP AAPSSGMHDN GARCHAVEIE AKILGLEAMI 300 RVQCHKRNHP AAKLMTALRE LDLDVYHASV SVVKDIMIQQ VAVKMATRVY SQDQLNAALY 360 GRLAEPGTAM QIR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5gnj_A | 1e-25 | 198 | 265 | 10 | 77 | Transcription factor MYC2 |
| 5gnj_B | 1e-25 | 198 | 265 | 10 | 77 | Transcription factor MYC2 |
| 5gnj_E | 1e-25 | 198 | 265 | 10 | 77 | Transcription factor MYC2 |
| 5gnj_F | 1e-25 | 198 | 265 | 10 | 77 | Transcription factor MYC2 |
| 5gnj_G | 1e-25 | 198 | 265 | 10 | 77 | Transcription factor MYC2 |
| 5gnj_I | 1e-25 | 198 | 265 | 10 | 77 | Transcription factor MYC2 |
| 5gnj_M | 1e-25 | 198 | 265 | 10 | 77 | Transcription factor MYC2 |
| 5gnj_N | 1e-25 | 198 | 265 | 10 | 77 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator involved in jasmonate (JA) signaling pathway during spikelet development. Binds to the G2 region G-box (5'-CACGTG-3') of the MADS1 promoter and thus directly regulates the expression of MADS1. Its function in MADS1 activation is abolished by TIFY3/JAZ1 which directly target MYC2 during spikelet development. {ECO:0000269|PubMed:24647160}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00086 | PBM | Transfer from AT4G17880 | Download |
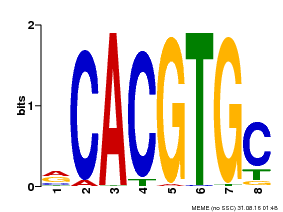
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK354796 | 0.0 | AK354796.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv1011F06. | |||
| GenBank | AK357816 | 0.0 | AK357816.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv1062G16. | |||
| GenBank | AK359813 | 0.0 | AK359813.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv1103K10. | |||
| GenBank | AK360465 | 0.0 | AK360465.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv1118K08. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020165313.1 | 1e-115 | transcription factor MYC2-like | ||||
| Swissprot | Q336P5 | 1e-99 | MYC2_ORYSJ; Transcription factor MYC2 | ||||
| TrEMBL | M7Z8G6 | 0.0 | M7Z8G6_TRIUA; Transcription factor MYC4 | ||||
| STRING | TRIUR3_32014-P1 | 0.0 | (Triticum urartu) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP4452 | 31 | 57 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G17880.1 | 2e-60 | bHLH family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



