 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | TRIUR3_30534-P1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Triticum
|
||||||||
| Family | TALE | ||||||||
| Protein Properties | Length: 465aa MW: 52878.8 Da PI: 9.2253 | ||||||||
| Description | TALE family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 26.9 | 8.1e-09 | 207 | 245 | 17 | 55 |
HHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHH CS
Homeobox 17 elFeknrypsaeereeLAkklgLterqVkvWFqNrRake 55
e+ +k +yps++++ LA+++gL+ +q+ +WF N+R ++
TRIUR3_30534-P1 207 EMHYKWPYPSESQKVALAESTGLDLKQINNWFINQRKRH 245
5566789*****************************885 PP
| |||||||
| 2 | ELK | 36.7 | 9.1e-13 | 165 | 186 | 1 | 22 |
ELK 1 ELKhqLlrKYsgyLgsLkqEFs 22
ELKh+Ll+KYsgyL+sLkqE+s
TRIUR3_30534-P1 165 ELKHHLLKKYSGYLSSLKQELS 186
9*******************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01256 | 1.3E-28 | 58 | 109 | IPR005541 | KNOX2 |
| Pfam | PF03791 | 6.0E-23 | 64 | 107 | IPR005541 | KNOX2 |
| SMART | SM01188 | 1800 | 67 | 87 | IPR005539 | ELK domain |
| Pfam | PF03789 | 1.1E-9 | 165 | 186 | IPR005539 | ELK domain |
| SMART | SM01188 | 4.9E-7 | 165 | 186 | IPR005539 | ELK domain |
| PROSITE profile | PS51213 | 11.331 | 165 | 185 | IPR005539 | ELK domain |
| PROSITE profile | PS50071 | 12.665 | 185 | 248 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 4.06E-19 | 186 | 261 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 6.1E-13 | 187 | 252 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 1.5E-27 | 190 | 249 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 6.13E-11 | 197 | 249 | No hit | No description |
| Pfam | PF05920 | 2.2E-16 | 205 | 244 | IPR008422 | Homeobox KN domain |
| PROSITE pattern | PS00027 | 0 | 223 | 246 | IPR017970 | Homeobox, conserved site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0001708 | Biological Process | cell fate specification | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010051 | Biological Process | xylem and phloem pattern formation | ||||
| GO:0010089 | Biological Process | xylem development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 465 aa Download sequence Send to blast |
MEEIGHHFGL GATAHGQHHS QLPWGSSPLS AVGAPPEVLA RLTAVAQDLE LRQRTALGSL 60 GTATEPELDQ FMEAYHEMLV KYREELTRPL QEAMEFLRRV ETQLNSLSIS GRSLRNILSS 120 GKAKRDTENL PGSKKQPKEN KREDQEGSGG ETELPEIDAH GVDQELKHHL LKKYSGYLSS 180 LKQELSKKKK KGKLPKDARQ QLLSWWEMHY KWPYPSESQK VALAESTGLD LKQINNWFIN 240 QRKRHWKPSD EMQFVMMDAY HPPNAAFYMD GHFVNDSGLY SAPVTHMLST ADALVPRHPP 300 FLPLFSPFPV QPWPSAFQAI ETRLLYEAEY PVSSNMWVPG AWRISAGGVP VPTLPTGAAR 360 RAEIACIRVN LPQAAREGPR DVHDSPLWEP YFRRRHAEQL EATNDVVPSR RLNSEGRRRW 420 WGVPDRTLEA ILEYIEGGNT PRLEYPAPPS FSRRRGSSWP PRYAR |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | May play a role in meristem formation and/or maintenance. Overexpression causes the hooded phenotype characterized by the appearance of an extra flower of inverse polarity on the lemma. Binds to the DNA sequence 5'-TGAC-3'. | |||||
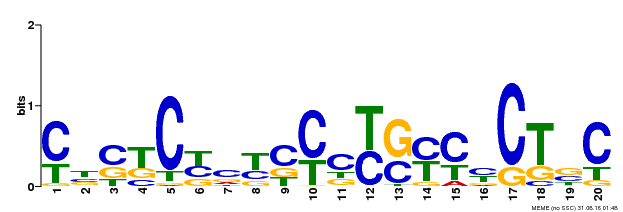
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00675 | ChIP-seq | Transfer from GRMZM2G017087 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF224498 | 0.0 | AF224498.1 Triticum aestivum KNOTTED-1-like homeobox protein a (knox1a) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020147582.1 | 1e-151 | homeobox protein KNOX3 | ||||
| Swissprot | Q43484 | 1e-149 | KNOX3_HORVU; Homeobox protein KNOX3 | ||||
| TrEMBL | M8AF24 | 0.0 | M8AF24_TRIUA; Homeobox protein KNOX3 | ||||
| STRING | TRIUR3_30534-P1 | 0.0 | (Triticum urartu) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP9886 | 31 | 45 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G08150.1 | 7e-84 | KNOTTED-like from Arabidopsis thaliana | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



