 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | TRIUR3_29784-P1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Triticum
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 293aa MW: 32028.1 Da PI: 4.9129 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 47.4 | 4.9e-15 | 155 | 200 | 4 | 52 |
AP2 4 kGVrwdkkrgrWvAeIrdpsengkr.krfslgkfgtaeeAakaaiaarkk 52
GVr+++ +g+++AeIrd + + r++lg+f+t+e Aa +++a+
TRIUR3_29784-P1 155 IGVRRRP-WGKYAAEIRDSTR---SgERVWLGTFDTPEAAALTYDQAAYT 200
6******.**********322...35*********************975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51032 | 20.929 | 153 | 211 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 5.3E-7 | 154 | 165 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 5.17E-20 | 154 | 213 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 3.6E-10 | 155 | 200 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 1.8E-26 | 156 | 212 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 7.06E-22 | 156 | 212 | No hit | No description |
| SMART | SM00380 | 4.8E-27 | 156 | 217 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 5.3E-7 | 177 | 193 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0006952 | Biological Process | defense response | ||||
| GO:0009873 | Biological Process | ethylene-activated signaling pathway | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 293 aa Download sequence Send to blast |
MEPNHNGYGY YDHNNGSYTN DDNPCYYGMS YASSEPSSYG YQYQQSPSTQ PQSSYDNHYY 60 AAYDITNSSS SSAAPSYAAA DDIALYAQQQ HPQYLNFGGG SYQYDGAASA MGMGMGMDMD 120 QFSALMEATS ISTAGPSWAE QQEAKKAKAE PPQLIGVRRR PWGKYAAEIR DSTRSGERVW 180 LGTFDTPEAA ALTYDQAAYT SRGPAAVLNF PVERVRESLR VLKLGKAAAA GEDDSPVLAL 240 KRRHCMRKRT PKGKGQEGDV KKKKQPAADT ASSVLELEDL GADYLEELLA LSE |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5wx9_A | 6e-26 | 142 | 213 | 3 | 74 | Ethylene-responsive transcription factor ERF096 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 246 | 263 | RKRTPKGKGQEGDVKKKK |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00374 | DAP | Transfer from AT3G23240 | Download |
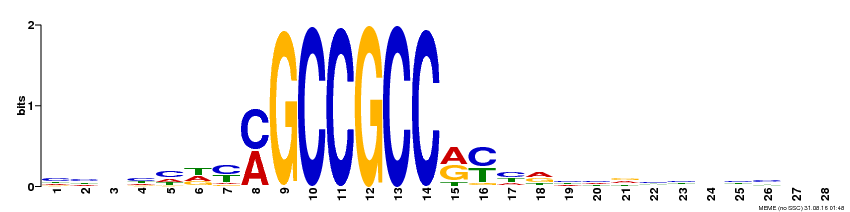
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC130602 | 4e-33 | AC130602.5 Oryza sativa Japonica Group cultivar Nipponbare chromosome 5 clone B1122D01, complete sequence. | |||
| GenBank | AP014961 | 4e-33 | AP014961.1 Oryza sativa Japonica Group DNA, chromosome 5, cultivar: Nipponbare, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020196844.1 | 1e-166 | ethylene-responsive transcription factor 15-like | ||||
| TrEMBL | T1NGN2 | 0.0 | T1NGN2_TRIUA; Uncharacterized protein | ||||
| STRING | TRIUR3_29784-P1 | 0.0 | (Triticum urartu) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP277 | 37 | 249 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G23240.1 | 1e-42 | ethylene response factor 1 | ||||



