 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | TRIUR3_28076-P1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Triticum
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 295aa MW: 32561.8 Da PI: 6.7151 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 21.7 | 4.6e-07 | 110 | 155 | 2 | 45 |
SSS-HHHHHHHHHHHHHTTTT...-HHHHHHHHTTTS-HHHHHHHHH CS
Myb_DNA-binding 2 grWTteEdellvdavkqlGgg...tWktIartmgkgRtlkqcksrwq 45
g+WT eE + + +a + + +W+ +a +++ ++t +++++++
TRIUR3_28076-P1 110 GAWTLEENKVFEEALAAIDLDapdRWEMVALMLP-RKTVADVVNHYR 155
79*****************99*************.***********8 PP
| |||||||
| 2 | Myb_DNA-binding | 33.6 | 9e-11 | 217 | 253 | 3 | 39 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkq 39
+WT+eE+ l++++ k++G g+W+ I+r + Rt+ q
TRIUR3_28076-P1 217 PWTEEEHRLFLKGLKKYGRGDWRNISRNYVTSRTPTQ 253
8***************************999999865 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50090 | 6.11 | 104 | 158 | IPR017877 | Myb-like domain |
| SMART | SM00717 | 8.2E-6 | 108 | 160 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 8.61E-10 | 110 | 164 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 6.77E-5 | 111 | 157 | No hit | No description |
| PROSITE profile | PS51294 | 10.797 | 210 | 256 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.58E-9 | 211 | 253 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 4.2E-9 | 213 | 253 | IPR006447 | Myb domain, plants |
| Gene3D | G3DSA:1.10.10.60 | 6.8E-11 | 214 | 253 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 0.0022 | 214 | 256 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.05E-9 | 217 | 253 | No hit | No description |
| Pfam | PF00249 | 3.2E-8 | 217 | 253 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 295 aa Download sequence Send to blast |
MVFFKDISSG SSSGDYSFTT RIQDPPYRHC NRLAEQEHVI TSAPKRASFH LPPSPLSCLP 60 PLLAVPKPAQ VAATRADMMA EAWADVLPPP APYFAGQPCW AVQPERRGGG AWTLEENKVF 120 EEALAAIDLD APDRWEMVAL MLPRKTVADV VNHYRALEND VGFIEAGLVP FPHYDSSSPS 180 SGFTLDWDGG AAGAGGFRRG YCLKRGRADQ ERKKGVPWTE EEHRLFLKGL KKYGRGDWRN 240 ISRNYVTSRT PTQPGSLGGG HPYGSVKLEH QSSFMDGMVV DLDESLLLQM QCGQL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2cjj_A | 3e-14 | 106 | 177 | 5 | 76 | RADIALIS |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Involved in the dorsovental asymmetry of flowers. Promotes ventral identity. {ECO:0000269|PubMed:11937495, ECO:0000269|PubMed:9118809}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
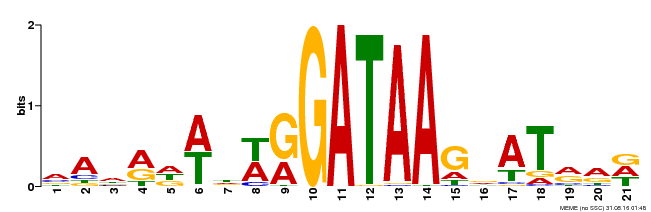
| Motif ID | Method | Source | Motif file |
| MP00296 | DAP | Transfer from AT2G38090 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK361442 | 0.0 | AK361442.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv1140K03. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020182102.1 | 1e-119 | transcription factor DIVARICATA-like | ||||
| Swissprot | Q8S9H7 | 7e-52 | DIV_ANTMA; Transcription factor DIVARICATA | ||||
| TrEMBL | M7ZMD0 | 0.0 | M7ZMD0_TRIUA; Transcription factor MYB1R1 | ||||
| STRING | TRIUR3_28076-P1 | 0.0 | (Triticum urartu) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2274 | 37 | 96 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G58900.1 | 2e-55 | Homeodomain-like transcriptional regulator | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



