 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | TRIUR3_27886-P1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Triticum
|
||||||||
| Family | RAV | ||||||||
| Protein Properties | Length: 305aa MW: 33195.8 Da PI: 6.6423 | ||||||||
| Description | RAV family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 39.8 | 1.1e-12 | 39 | 87 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s++kGV + +grW A+I++ + r++lg+f +e Aa+a++ a+++++g
TRIUR3_27886-P1 39 SRFKGVVPQP-NGRWGAQIYEK-----HSRVWLGTFADEESAARAYDVAALRFRG 87
689***9788.8*********3.....5*************************98 PP
| |||||||
| 2 | B3 | 95.2 | 4.6e-30 | 158 | 250 | 1 | 85 |
EEEE-..-HHHHTT-EE--HHH.HTT........---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE- CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh........ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkld 85
f+k+ tpsdv+k++rlv+pk++ae+h g++ ++ l +ed +g++W+++++y+++s++yvltkGW++Fv+++gL +gD+v+F+++
TRIUR3_27886-P1 158 FEKAVTPSDVGKLNRLVVPKQHAEKHfppttaaaTGSNGKGLLLNFEDGEGKVWRFRYSYWNSSQSYVLTKGWSRFVREKGLVAGDTVTFSRS 250
89****************************999955667899************************************************943 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF54171 | 1.83E-16 | 39 | 96 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 2.0E-7 | 39 | 87 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 20.389 | 40 | 95 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 3.71E-22 | 40 | 95 | No hit | No description |
| SMART | SM00380 | 9.0E-27 | 40 | 101 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 4.2E-20 | 40 | 95 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:2.40.330.10 | 7.5E-38 | 154 | 268 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 3.80E-28 | 156 | 250 | No hit | No description |
| SuperFamily | SSF101936 | 2.35E-29 | 156 | 250 | IPR015300 | DNA-binding pseudobarrel domain |
| Pfam | PF02362 | 1.2E-27 | 158 | 252 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 13.86 | 158 | 269 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 4.6E-21 | 158 | 258 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 305 aa Download sequence Send to blast |
MASGKPTNPE IDNDMECSSP ESGAEDAVES SSPAPAPSSR FKGVVPQPNG RWGAQIYEKH 60 SRVWLGTFAD EESAARAYDV AALRFRGRDA VTNYQLPLAV EGASSSSTSE LAFLADHSKA 120 EIVDMLRKHT YADELRQGLR RGHGRAQPTP AWAREFLFEK AVTPSDVGKL NRLVVPKQHA 180 EKHFPPTTAA ATGSNGKGLL LNFEDGEGKV WRFRYSYWNS SQSYVLTKGW SRFVREKGLV 240 AGDTVTFSRS AYVMNDTEEQ LFIDYKQKNK NDEAADAAKN EAGHAAVKLF GVDIGGGEMA 300 GSSGG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wid_A | 7e-49 | 155 | 272 | 11 | 121 | DNA-binding protein RAV1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00025 | PBM | Transfer from AT1G68840 | Download |
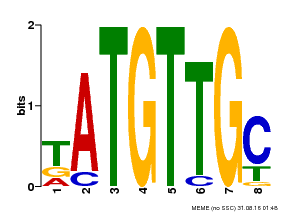
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK361289 | 0.0 | AK361289.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv1136P02. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020164924.1 | 0.0 | AP2/ERF and B3 domain-containing protein Os01g0141000-like | ||||
| Swissprot | Q9AWS7 | 1e-122 | Y1407_ORYSJ; Putative AP2/ERF and B3 domain-containing protein Os01g0140700 | ||||
| TrEMBL | T1NBI8 | 0.0 | T1NBI8_TRIUA; Uncharacterized protein | ||||
| STRING | TRIUR3_27886-P1 | 0.0 | (Triticum urartu) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP456 | 38 | 203 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G68840.2 | 1e-101 | related to ABI3/VP1 2 | ||||



