 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | TRIUR3_25447-P1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Triticum
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 291aa MW: 31347 Da PI: 8.512 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 52.8 | 9.3e-17 | 8 | 53 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
rg+W++eEd l +v+++G ++W++I r ++ gR++k+c++rw +
TRIUR3_25447-P1 8 RGPWSPEEDGALRLLVERHGARNWTAIGRGVP-GRSGKSCRLRWCNQ 53
89******************************.***********985 PP
| |||||||
| 2 | Myb_DNA-binding | 51.6 | 2.1e-16 | 60 | 102 | 1 | 45 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwq 45
r ++T+eEd +++a+++lG++ W++Iar + gRt++ +k++w+
TRIUR3_25447-P1 60 RRPFTAEEDAAIARAHARLGNR-WAAIARLLR-GRTDNAVKNHWN 102
679*******************.*********.***********8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 23.486 | 1 | 58 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.28E-29 | 5 | 101 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 8.1E-14 | 7 | 56 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 4.7E-16 | 8 | 53 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 9.5E-23 | 9 | 60 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.73E-13 | 10 | 52 | No hit | No description |
| SMART | SM00717 | 1.3E-14 | 59 | 107 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS51294 | 17.501 | 60 | 109 | IPR017930 | Myb domain |
| Pfam | PF00249 | 4.9E-14 | 60 | 102 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.7E-21 | 61 | 109 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 2.41E-11 | 62 | 102 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:2000022 | Biological Process | regulation of jasmonic acid mediated signaling pathway | ||||
| GO:2000031 | Biological Process | regulation of salicylic acid mediated signaling pathway | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 291 aa Download sequence Send to blast |
MEGCDRIRGP WSPEEDGALR LLVERHGARN WTAIGRGVPG RSGKSCRLRW CNQLSPGVER 60 RPFTAEEDAA IARAHARLGN RWAAIARLLR GRTDNAVKNH WNCSLKRRLA DVGVAEEERP 120 GKRGGVTPES TSGSGSGSGS GSDRSDLSHR SVFGLGQVYR PVARAGGFEP ADCVMSRRHE 180 VEREEAEDPL TSLSLSLPGT ATEVQGFHHD SSHSHLHQPS PSSPSPSPPP APTSYPFSPE 240 FAAMMQQMIR DEVCRYISGV GCGANLPSMP QVVDGVMRAA AERAGGVART Q |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 7e-36 | 5 | 109 | 4 | 108 | B-MYB |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00596 | DAP | Transfer from AT5G67300 | Download |
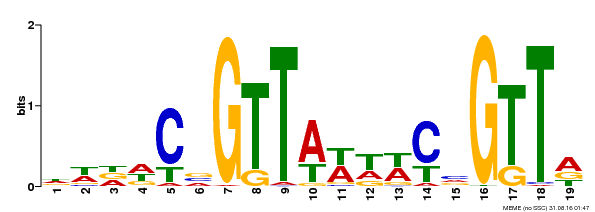
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK364159 | 1e-153 | AK364159.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv2022D22. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020162234.1 | 1e-148 | transcription factor MYB44-like | ||||
| TrEMBL | M8A613 | 0.0 | M8A613_TRIUA; Transcription factor MYB44 | ||||
| STRING | TRIUR3_25447-P1 | 0.0 | (Triticum urartu) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP789 | 38 | 154 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G50060.1 | 4e-46 | myb domain protein 77 | ||||



